DNA
Int J Radiat Biol. 2011 Apr;87(4):409-15. doi: 10.3109/09553002.2011.538130. Epub 2011 Feb 28.
DNA is a fractal antenna in electromagnetic fields.
Blank M, Goodman R.Source
Department of Physiology, Columbia University, New York 10032, USA. [email protected]
Abstract
PURPOSE: To review the responses of deoxyribonucleic acid (DNA) to electromagnetic fields (EMF) in different frequency ranges, and characterise the properties of DNA as an antenna.
MATERIALS AND METHODS: We examined published reports of increased stress protein levels and DNA strand breaks due to EMF interactions, both of which are indicative of DNA damage. We also considered antenna properties such as electronic conduction within DNA and its compact structure in the nucleus.
RESULTS: EMF interactions with DNA are similar over a range of non-ionising frequencies, i.e., extremely low frequency (ELF) and radio frequency (RF) ranges. There are similar effects in the ionising range, but the reactions are more complex.
CONCLUSIONS: The wide frequency range of interaction with EMF is the functional characteristic of a fractal antenna, and DNA appears to possess the two structural characteristics of fractal antennas, electronic conduction and self symmetry. These properties contribute to greater reactivity of DNA with EMF in the environment, and the DNA damage could account for increases in cancer epidemiology, as well as variations in the rate of chemical evolution in early geologic history.
Comment in
DNA is a fractal antenna in electromagnetic fields.
Blank M, Goodman R.Source
Department of Physiology, Columbia University, New York 10032, USA. [email protected]
Abstract
PURPOSE: To review the responses of deoxyribonucleic acid (DNA) to electromagnetic fields (EMF) in different frequency ranges, and characterise the properties of DNA as an antenna.
MATERIALS AND METHODS: We examined published reports of increased stress protein levels and DNA strand breaks due to EMF interactions, both of which are indicative of DNA damage. We also considered antenna properties such as electronic conduction within DNA and its compact structure in the nucleus.
RESULTS: EMF interactions with DNA are similar over a range of non-ionising frequencies, i.e., extremely low frequency (ELF) and radio frequency (RF) ranges. There are similar effects in the ionising range, but the reactions are more complex.
CONCLUSIONS: The wide frequency range of interaction with EMF is the functional characteristic of a fractal antenna, and DNA appears to possess the two structural characteristics of fractal antennas, electronic conduction and self symmetry. These properties contribute to greater reactivity of DNA with EMF in the environment, and the DNA damage could account for increases in cancer epidemiology, as well as variations in the rate of chemical evolution in early geologic history.
Comment in
- Comments on DNA as a fractal antenna. [Int J Radiat Biol. 2011]
Regarding Wave Genetics, Andras Laszlo, June 2014
we may have had an exchange about this before. first of all there are no such studies as stated at the beginning of the article...that statement came from a journalist who translated what he was told. The important thing about wave genetics is a few sentences down.."junk DNA". This term was coined by Sushu Ohno from City of Hope, as a tongue in cheek term at a meeting in 1972, when our understanding of DNA organization was truly elementary. If you google "junk DNA", you will see that in 2012, 31 papers that came out in August, with people from all over the world, have finally put the silver or gold nail in the coffin of this concept, which says that a lot of our DNA is not important. This is just not true. There is a lot of non-protein coding DNA sequences that are made into lots of different RNAs. micro RNAs, long repeat RNAs, which are have been shown everything from cancer to stem cells. Given that the basic premise of wave genetics rests on a concept that has been thoroughly disproven, the wave genetics edifice collapses. The difficulty is that Gariaev is a very smart man, but he is a physicist and biphysicist, and has very little feeling for biology. All of his published studies have been made with purified DNA in the test tube. THe difficulty is that DNA as a polymer is very complex and has some fascinating properties which have nothing to do with its biological function. Thus one cannot extrapolate from experiments with purified DNA. Inside cells, even the simplest bacteria, DNA is always associated with proteins and packed very tightly. Our cells contain about 2 meters of DNA, which needs to be packed into a nucleus of 5-10 micron diameter. Very different situation from experiments performed with dilute solutions....hope this makes sense....
we may have had an exchange about this before. first of all there are no such studies as stated at the beginning of the article...that statement came from a journalist who translated what he was told. The important thing about wave genetics is a few sentences down.."junk DNA". This term was coined by Sushu Ohno from City of Hope, as a tongue in cheek term at a meeting in 1972, when our understanding of DNA organization was truly elementary. If you google "junk DNA", you will see that in 2012, 31 papers that came out in August, with people from all over the world, have finally put the silver or gold nail in the coffin of this concept, which says that a lot of our DNA is not important. This is just not true. There is a lot of non-protein coding DNA sequences that are made into lots of different RNAs. micro RNAs, long repeat RNAs, which are have been shown everything from cancer to stem cells. Given that the basic premise of wave genetics rests on a concept that has been thoroughly disproven, the wave genetics edifice collapses. The difficulty is that Gariaev is a very smart man, but he is a physicist and biphysicist, and has very little feeling for biology. All of his published studies have been made with purified DNA in the test tube. THe difficulty is that DNA as a polymer is very complex and has some fascinating properties which have nothing to do with its biological function. Thus one cannot extrapolate from experiments with purified DNA. Inside cells, even the simplest bacteria, DNA is always associated with proteins and packed very tightly. Our cells contain about 2 meters of DNA, which needs to be packed into a nucleus of 5-10 micron diameter. Very different situation from experiments performed with dilute solutions....hope this makes sense....
Networks of Genes Respond to Social Experiences October 13, 2013 It is extremely surprising how networks of hundreds of genes respond immediately to human interactions and thoughts—despite the fact that actions of humans are eight orders of magnitude larger than molecular genetic events. But, it is, perhaps, more remarkable that networks of genes respond rapidly to social experiences. Previous posts have discussed the immediate neuroplasticity that occurs in widespread circuits with very complex detailed genetic production of new proteins, including motors, tubules, receptors, and neurotransmitters. The immune system does the same with cytokines, receptors, and antibodies. Now, it has become clear that meditation, tai chi, social interactions, abuse, charitable actions all affect very specific networks of genes in the nervous system and the immune system.
But, how can a network of genes deep inside the nucleus respond to events as if it is a brain? In fact, we don’t know how brains respond in such a detailed genetic manner, but having the genes of individual cells, deep in the nucleus respond as if it is a brain is truly remarkable. When networks of genes respond to social experience, it is further evidence of mind interacting at all levels at once, including the 12 orders of magnitude from world civilizations, to molecules in the nucleus.
Previous posts show how dramatic changes occur in neuronal genes instantly with thought or behavior with neuroplasticity. Recently, it was shown that exercise effects 10,000 genes, that insulin effects 7 thousand genes in the cell. Meditation, a conscious mental activity, causes changes in thousands of genes in immune cells.
Human action or thought alters genes by expressing certain ones and suppressing others. There appears to be a dimmer switch on each gene that can increase or decrease activity based upon many levels of regulation (methylation of DNA, methylation of histones, regulatory molecules such as promoters and inhibitors). The process is even more complex through alternative RNA editing of the code. These alterations of RNA through cells’ self editing of messenger RNA create very particular shaped proteins to express a particular thought, feeling, or activity.
Bees In New Environment Modifies Large Network of Genes Changing environments dramatically alters genes, almost immediately. In a study, infant bees were taken from two very different colonies—one group from the mild mannered European bees and another from the very aggressive African bees. They were then switched and put in the opposite tribe’s hive. Growing up in the new hives, the African bees became calm and the European bees became aggressive.
Remarkably, as the bees changed in their new homes, large networks of genes were completely altered. Being in the new environment completely remapped the networks of genes in a short time. As the bees entirely changed their personality and behavior, they looked more and more like the bees in the new hive. And their genes’ activity looked like them, also.
Every Cell Has All the Genes Every cell has all genes available; so, the difference between stem cells, kidney cells and brain cells is expression of sets of genes in different patterns. Changes in genetic expression occur in specially timed sequences in the fetus, childhood, adolescence and adulthood.
Gene changes, also, occur based on environment, both long term and short term. Most remarkably, they change based upon behavior and thought. Genes switch on to fight different infections, or can become unhinged and develop cancer. If too many genes change, the entire nature of the cell could be different; the entire nature of an organism could be different.
Danger Signals With signals of danger, such as sounds or smells, large numbers of genes become activated at the same time in large networks. The greater the aggression, the greater are the genetic changes. The genes and behavior change together. Hearing a pleasant signal or a danger signal immediately activates and dampens completely different genetic networks. In animal research these gene changes occurred within minutes.
Observing change in gene networks revealed critical specific regulatory genes that are responsible for altering a large group of other genes. These “immediate early genes” were previously known to occur in immunity and in eating. Recently, these same regulatory genes have been found to change entire networks of genes based upon social and behavior experiences.
In one experiment, a dominant fish was removed from a tank and almost immediately the second ranked fish had massive changes in large networks of genes within several hours. In fact, this fish grew 20% in several hours.
Epigenetic Changes Previous posts have described the dramatic findings of ENCODE (Encyclopedia of DNA Elements research consortium of 160 international research centers), which showed that millions of regulatory RNA particles are produced by DNA as well as proteins. These regulatory particles, (including microRNA, long and short non coding RNA, and protein promoters) have dramatic effects on DNA. (See posts on ENCODE and RNA regulatory particles.)
Another type of DNA regulation is epigenetic marking, including methylation of DNA and histones.
Epigenetic markings effect changes that can be inherited for generations. Mothers exposed to famine had altered genes affecting their health. The alterations, also, affected their children and grandchildren. Mothering and caring behavior can change the epigenetic markings and create different genetic network expressions.
Stress related genes alter methylation of DNA, but long-term behavioral changes occur because of neural circuits. How does methylation translate into neural circuits?
Social Experiences Have Powerful Effects on Genes The environment influences the organism as new cells are built each day. Billions of new blood cells, skin cells, and mucosal cells are made each day and these cells’ behavior are triggered by changing DNA networks in response to our experiences, and the chemical environment we are passing through. In the brain many new glial cells and a small number of neurons are, also, made each day.
Social connectivity has powerful effects on genes. The gene networks of social experience are consistent through many animals.
Social isolation is much more devastating than stress—the best-known disease risk factor. Isolated poor people do very poorly, while high pressure stressed people in good networks do well. The diseases of isolation are obesity, diabetes, hypertension, coronary heart disease, and strokes.
Feeling close to others (even if they are not physically present for isolated or imprisoned people) will protect the body with positive gene changes. Experience is what we take from the environment. Perhaps, this is one reason a spiritual teacher provides such support. The fact that subjective perception strongly affects the immune system is evidence of the power of the mind.
Previous posts described the details of dramatic immune changes with both meditation and charitable service. (See post Meditation Update for more details). Both of these are conscious behaviors.
With pleasure from charitable community service (not pleasure from other self oriented activities) there is a decrease in the gene activity of pro-inflammatory cytokines such as IL1B, IL6, IL8, and TNF. There was increased expression of genes involved in type I IFN antiviral responses including IFI-, OAS-, and MX- family genes. There was also increased IgG1 antibody synthesis.
With meditation, yoga, Tai Chi and other practices many positive immune changes occur. These include decreased immune inflammatory factors interleukin 6, and NF-KappaB, and an increase in the important antiviral factor IRF1. Other studies showed decreased inflammation with local skin burns, fewer colds and decreased stress hormones.
Long term meditators and novices both showed epigenetic gene expression changes related to increased mitochondrial resilience. The genes that changed related to very significant functions including energy metabolism, mitochondrial function, insulin secretion, telomere maintenance and decrease in inflammation and oxidative stress responses. The meditators had less respiratory infections. Meditating dementia caregivers had 68 gene changes related to decreased inflammation.
This demonstrates that conscious mental activities and behaviors can have major effects on our immune systems.
Genes and Social Behavior The social world outside determines what the genes do within the nucleus of the cell.
Social experiences can impact genes in many different ways. Genes have regions that if triggered by stress will release cortisol. Another region will stimulate norepinephrine and dopamine to trigger the body’s fight and flight response in cells throughout many organs. These two triggers exist in various places in the genes and can create a variety of different proteins.
The brain responds to social situations by stimulating hormones, immune cytokines and neurotransmitters to produce transcription factors that will alter gene networks. The hypothalamic-pituitary-adrenal (HPA) and sympathetic systems are powerful gene activators. Signaling molecules trigger receptors on the cell surface, then a cascade to the nucleus stimulates the genes. Different transcription factors produce different pathways, such as Nf-kB and CREB.
These different gene networks form a wiring diagram of genetic response. The entire normal brain response to ordinary signals can be altered in this process.
In the isolation experience, for example, the factor NF-kappaB that drives inflammation becomes very important in determining the specific response of signals. Cortisol, which normally inhibits NF-kB, doesn’t do this under stress and isolation. It does the opposite. Therefore, the response to the entire HPA signaling is altered.
The brain is signaling to decrease inflammation, but the receptors and the cascades ignore this. Isolation and social loss both disconnect a critical normal physiological mechanism. This is just one example of complex genetic mechanisms that respond to abuse, isolation, and other social circumstances.
The response to social situations, also, alters RNA editing and transcription, changing the entire network of genetic signals, just as if it were a brain with a new circuit. Chronic stress increases a factor, NGF, and this increases sympathetic nerves in the lymph nodes (the brain circuits of the immune system). As this nerve changes in the lymph node, the response to a virus is decreased. The entire relationship between the immune and nervous system has shifted. Later, interferon genes are inhibited in this process, which changes future responses as well.
In this way an experience creates a new circuit in the immune/nervous genetic systems, which can last for years.
Subjective Mind Changes Genes – Not Just External Situations The place of the mind becomes clearer in this analysis. The psychological perceptions and experiences become the way genetic circuits are modeled—by the perceptions of external events, not the events themselves. It is the subjective mental awareness of these events that determines the genetic rewiring.
Events can alter extensive wiring diagrams through specific genetic pathways. Mind changes how the brain uses its circuits. And mind changes the ways that genetic circuits in cells are altered. In brains measuring MRI doesn’t tell mechanisms or cause and effect. In the same way, measuring genetic circuits doesn’t tell cause and effect.
Networks of Genes Respond to Social Experiences Where is the Brain in the Gene?
It is quite remarkable that brains are able to respond to situations and mental events with almost instantaneous changes in wide ranging circuits, including many different very complex molecular changes in different neurons and astrocytes. A second later, a different circuit of neurons, some including the same neurons and some not, suddenly respond to the next event.
But, it is far more extraordinary to consider that social situations almost instantly trigger networks of genes, deep inside the cell. It appears that there are specific genetic hubs that can suddenly trigger thousands of genes in different ways–stimulating some and inhibiting others. These events trigger far reaching networks of cells, all at once, in the immune system, the hormonal systems, bodily organs and the nervous system. What is the brain deep inside the cell’s nucleus where networks of genes respond to social situations?
It is subjective mind and perception that changes genes, not just external situations. This is further evidence of mind affecting a large number of orders of magnitude simultaneously.
Copyright Jon Lieff 2011. - See more at: http://jonlieffmd.com/blog/networks-of-genes-respond-to-social-experiences#sthash.ZrubKwMy.X1eUL7Y9.dpuf
But, how can a network of genes deep inside the nucleus respond to events as if it is a brain? In fact, we don’t know how brains respond in such a detailed genetic manner, but having the genes of individual cells, deep in the nucleus respond as if it is a brain is truly remarkable. When networks of genes respond to social experience, it is further evidence of mind interacting at all levels at once, including the 12 orders of magnitude from world civilizations, to molecules in the nucleus.
Previous posts show how dramatic changes occur in neuronal genes instantly with thought or behavior with neuroplasticity. Recently, it was shown that exercise effects 10,000 genes, that insulin effects 7 thousand genes in the cell. Meditation, a conscious mental activity, causes changes in thousands of genes in immune cells.
Human action or thought alters genes by expressing certain ones and suppressing others. There appears to be a dimmer switch on each gene that can increase or decrease activity based upon many levels of regulation (methylation of DNA, methylation of histones, regulatory molecules such as promoters and inhibitors). The process is even more complex through alternative RNA editing of the code. These alterations of RNA through cells’ self editing of messenger RNA create very particular shaped proteins to express a particular thought, feeling, or activity.
Bees In New Environment Modifies Large Network of Genes Changing environments dramatically alters genes, almost immediately. In a study, infant bees were taken from two very different colonies—one group from the mild mannered European bees and another from the very aggressive African bees. They were then switched and put in the opposite tribe’s hive. Growing up in the new hives, the African bees became calm and the European bees became aggressive.
Remarkably, as the bees changed in their new homes, large networks of genes were completely altered. Being in the new environment completely remapped the networks of genes in a short time. As the bees entirely changed their personality and behavior, they looked more and more like the bees in the new hive. And their genes’ activity looked like them, also.
Every Cell Has All the Genes Every cell has all genes available; so, the difference between stem cells, kidney cells and brain cells is expression of sets of genes in different patterns. Changes in genetic expression occur in specially timed sequences in the fetus, childhood, adolescence and adulthood.
Gene changes, also, occur based on environment, both long term and short term. Most remarkably, they change based upon behavior and thought. Genes switch on to fight different infections, or can become unhinged and develop cancer. If too many genes change, the entire nature of the cell could be different; the entire nature of an organism could be different.
Danger Signals With signals of danger, such as sounds or smells, large numbers of genes become activated at the same time in large networks. The greater the aggression, the greater are the genetic changes. The genes and behavior change together. Hearing a pleasant signal or a danger signal immediately activates and dampens completely different genetic networks. In animal research these gene changes occurred within minutes.
Observing change in gene networks revealed critical specific regulatory genes that are responsible for altering a large group of other genes. These “immediate early genes” were previously known to occur in immunity and in eating. Recently, these same regulatory genes have been found to change entire networks of genes based upon social and behavior experiences.
In one experiment, a dominant fish was removed from a tank and almost immediately the second ranked fish had massive changes in large networks of genes within several hours. In fact, this fish grew 20% in several hours.
Epigenetic Changes Previous posts have described the dramatic findings of ENCODE (Encyclopedia of DNA Elements research consortium of 160 international research centers), which showed that millions of regulatory RNA particles are produced by DNA as well as proteins. These regulatory particles, (including microRNA, long and short non coding RNA, and protein promoters) have dramatic effects on DNA. (See posts on ENCODE and RNA regulatory particles.)
Another type of DNA regulation is epigenetic marking, including methylation of DNA and histones.
Epigenetic markings effect changes that can be inherited for generations. Mothers exposed to famine had altered genes affecting their health. The alterations, also, affected their children and grandchildren. Mothering and caring behavior can change the epigenetic markings and create different genetic network expressions.
Stress related genes alter methylation of DNA, but long-term behavioral changes occur because of neural circuits. How does methylation translate into neural circuits?
- Mothering sets up a calibration in the child as to how the brain responds to stress. More methyl groups with less nurturing mothers produce less receptors. Altered hormones affect the baby’s future behavior. Mice raised in groups are better socially and have more receptors. Stress activates more methylation of the gene of BDNF, which produces less BDNF affecting brain cells and neuroplasticity.
- Addiction is another example. It produced more acetylation of histones and decreased methylation of histones in the specific region of reward in the brain, nucleus accumbens, decreasing dendritic spines. More methylation also increased animals desire for cocaine.
- Another example is demonstrated in autopsies of suicide victims. They were abused as children and upon autopsy found to have more methyl groups on the cortisol gene.
Social Experiences Have Powerful Effects on Genes The environment influences the organism as new cells are built each day. Billions of new blood cells, skin cells, and mucosal cells are made each day and these cells’ behavior are triggered by changing DNA networks in response to our experiences, and the chemical environment we are passing through. In the brain many new glial cells and a small number of neurons are, also, made each day.
Social connectivity has powerful effects on genes. The gene networks of social experience are consistent through many animals.
- Studies show human loneliness predicted less immune response to microbes. HIV patients who were hiding their sexual orientation had greater amount of cancers and infections.
- People with more friends have fewer colds.
- Stress has major effects on the immune system. Monkeys with SIV (simian HIV) who were moved constantly into new social groups became ill more frequently. The immune system did not respond to the stress signal.
- Another study showed that lonely and engaged people had dramatic differences in hundreds their genes.
Social isolation is much more devastating than stress—the best-known disease risk factor. Isolated poor people do very poorly, while high pressure stressed people in good networks do well. The diseases of isolation are obesity, diabetes, hypertension, coronary heart disease, and strokes.
- Poor children showed more active inflammatory genes. Ambiguous social situations are threatening and affect immune genes. If the social scene is frightening then it affected gene networks, not just poverty.
- In abused children where negative gene changes occurred, those children who had one adult support experience monthly did not have this gene effect. The lack of connection was more damaging than the abuse. Isolation was the most damaging.
- With ovarian cancer 220 genes were activated for those women with less support and depression.
Feeling close to others (even if they are not physically present for isolated or imprisoned people) will protect the body with positive gene changes. Experience is what we take from the environment. Perhaps, this is one reason a spiritual teacher provides such support. The fact that subjective perception strongly affects the immune system is evidence of the power of the mind.
Previous posts described the details of dramatic immune changes with both meditation and charitable service. (See post Meditation Update for more details). Both of these are conscious behaviors.
With pleasure from charitable community service (not pleasure from other self oriented activities) there is a decrease in the gene activity of pro-inflammatory cytokines such as IL1B, IL6, IL8, and TNF. There was increased expression of genes involved in type I IFN antiviral responses including IFI-, OAS-, and MX- family genes. There was also increased IgG1 antibody synthesis.
With meditation, yoga, Tai Chi and other practices many positive immune changes occur. These include decreased immune inflammatory factors interleukin 6, and NF-KappaB, and an increase in the important antiviral factor IRF1. Other studies showed decreased inflammation with local skin burns, fewer colds and decreased stress hormones.
Long term meditators and novices both showed epigenetic gene expression changes related to increased mitochondrial resilience. The genes that changed related to very significant functions including energy metabolism, mitochondrial function, insulin secretion, telomere maintenance and decrease in inflammation and oxidative stress responses. The meditators had less respiratory infections. Meditating dementia caregivers had 68 gene changes related to decreased inflammation.
This demonstrates that conscious mental activities and behaviors can have major effects on our immune systems.
Genes and Social Behavior The social world outside determines what the genes do within the nucleus of the cell.
Social experiences can impact genes in many different ways. Genes have regions that if triggered by stress will release cortisol. Another region will stimulate norepinephrine and dopamine to trigger the body’s fight and flight response in cells throughout many organs. These two triggers exist in various places in the genes and can create a variety of different proteins.
The brain responds to social situations by stimulating hormones, immune cytokines and neurotransmitters to produce transcription factors that will alter gene networks. The hypothalamic-pituitary-adrenal (HPA) and sympathetic systems are powerful gene activators. Signaling molecules trigger receptors on the cell surface, then a cascade to the nucleus stimulates the genes. Different transcription factors produce different pathways, such as Nf-kB and CREB.
These different gene networks form a wiring diagram of genetic response. The entire normal brain response to ordinary signals can be altered in this process.
In the isolation experience, for example, the factor NF-kappaB that drives inflammation becomes very important in determining the specific response of signals. Cortisol, which normally inhibits NF-kB, doesn’t do this under stress and isolation. It does the opposite. Therefore, the response to the entire HPA signaling is altered.
The brain is signaling to decrease inflammation, but the receptors and the cascades ignore this. Isolation and social loss both disconnect a critical normal physiological mechanism. This is just one example of complex genetic mechanisms that respond to abuse, isolation, and other social circumstances.
The response to social situations, also, alters RNA editing and transcription, changing the entire network of genetic signals, just as if it were a brain with a new circuit. Chronic stress increases a factor, NGF, and this increases sympathetic nerves in the lymph nodes (the brain circuits of the immune system). As this nerve changes in the lymph node, the response to a virus is decreased. The entire relationship between the immune and nervous system has shifted. Later, interferon genes are inhibited in this process, which changes future responses as well.
In this way an experience creates a new circuit in the immune/nervous genetic systems, which can last for years.
Subjective Mind Changes Genes – Not Just External Situations The place of the mind becomes clearer in this analysis. The psychological perceptions and experiences become the way genetic circuits are modeled—by the perceptions of external events, not the events themselves. It is the subjective mental awareness of these events that determines the genetic rewiring.
Events can alter extensive wiring diagrams through specific genetic pathways. Mind changes how the brain uses its circuits. And mind changes the ways that genetic circuits in cells are altered. In brains measuring MRI doesn’t tell mechanisms or cause and effect. In the same way, measuring genetic circuits doesn’t tell cause and effect.
Networks of Genes Respond to Social Experiences Where is the Brain in the Gene?
It is quite remarkable that brains are able to respond to situations and mental events with almost instantaneous changes in wide ranging circuits, including many different very complex molecular changes in different neurons and astrocytes. A second later, a different circuit of neurons, some including the same neurons and some not, suddenly respond to the next event.
But, it is far more extraordinary to consider that social situations almost instantly trigger networks of genes, deep inside the cell. It appears that there are specific genetic hubs that can suddenly trigger thousands of genes in different ways–stimulating some and inhibiting others. These events trigger far reaching networks of cells, all at once, in the immune system, the hormonal systems, bodily organs and the nervous system. What is the brain deep inside the cell’s nucleus where networks of genes respond to social situations?
It is subjective mind and perception that changes genes, not just external situations. This is further evidence of mind affecting a large number of orders of magnitude simultaneously.
Copyright Jon Lieff 2011. - See more at: http://jonlieffmd.com/blog/networks-of-genes-respond-to-social-experiences#sthash.ZrubKwMy.X1eUL7Y9.dpuf
Feb. 2008
DNA Found to Have "Impossible" Telepathic Properties DNA has been found to have a bizarre ability to put itself together, even at a distance, when according to known science it shouldn't be able to. Explanation: None, at least not yet.
Scientists are reporting evidence that contrary to our current beliefs about what is possible, intact double-stranded DNA has the “amazing” ability to recognize similarities in other DNA strands from a distance. Somehow they are able to identify one another, and the tiny bits of genetic material tend to congregate with similar DNA. The recognition of similar sequences in DNA’s chemical subunits, occurs in a way unrecognized by science. There is no known reason why the DNA is able to combine the way it does, and from a current theoretical standpoint this feat should be chemically impossible.
Even so, the research published in ACS’ Journal of Physical Chemistry B, shows very clearly that homology recognition between sequences of several hundred nucleotides occurs without physical contact or presence of proteins. Double helixes of DNA can recognize matching molecules from a distance and then gather together, all seemingly without help from any other molecules or chemical signals.
In the study, scientists observed the behavior of fluorescently tagged DNA strands placed in water that contained no proteins or other material that could interfere with the experiment. Strands with identical nucleotide sequences were about twice as likely to gather together as DNA strands with different sequences. No one knows how individual DNA strands could possibly be communicating in this way, yet somehow they do. The “telepathic” effect is a source of wonder and amazement for scientists.
“Amazingly, the forces responsible for the sequence recognition can reach across more than one nanometer of water separating the surfaces of the nearest neighbor DNA,” said the authors Geoff S. Baldwin, Sergey Leikin, John M. Seddon, and Alexei A. Kornyshev and colleagues.
This recognition effect may help increase the accuracy and efficiency of the homologous recombination of genes, which is a process responsible for DNA repair, evolution, and genetic diversity. The new findings may also shed light on ways to avoid recombination errors, which are factors in cancer, aging, and other health issues.
http://www.dailygalaxy.com/my_weblog/2008/02/dna-found-to-ha.html
Scientists are reporting evidence that contrary to our current beliefs about what is possible, intact double-stranded DNA has the “amazing” ability to recognize similarities in other DNA strands from a distance. Somehow they are able to identify one another, and the tiny bits of genetic material tend to congregate with similar DNA. The recognition of similar sequences in DNA’s chemical subunits, occurs in a way unrecognized by science. There is no known reason why the DNA is able to combine the way it does, and from a current theoretical standpoint this feat should be chemically impossible.
Even so, the research published in ACS’ Journal of Physical Chemistry B, shows very clearly that homology recognition between sequences of several hundred nucleotides occurs without physical contact or presence of proteins. Double helixes of DNA can recognize matching molecules from a distance and then gather together, all seemingly without help from any other molecules or chemical signals.
In the study, scientists observed the behavior of fluorescently tagged DNA strands placed in water that contained no proteins or other material that could interfere with the experiment. Strands with identical nucleotide sequences were about twice as likely to gather together as DNA strands with different sequences. No one knows how individual DNA strands could possibly be communicating in this way, yet somehow they do. The “telepathic” effect is a source of wonder and amazement for scientists.
“Amazingly, the forces responsible for the sequence recognition can reach across more than one nanometer of water separating the surfaces of the nearest neighbor DNA,” said the authors Geoff S. Baldwin, Sergey Leikin, John M. Seddon, and Alexei A. Kornyshev and colleagues.
This recognition effect may help increase the accuracy and efficiency of the homologous recombination of genes, which is a process responsible for DNA repair, evolution, and genetic diversity. The new findings may also shed light on ways to avoid recombination errors, which are factors in cancer, aging, and other health issues.
http://www.dailygalaxy.com/my_weblog/2008/02/dna-found-to-ha.html
DNA Data Storage

: Scientists create rewritable digital data storage in DNA May 21, 2012
Scientists from Stanford's Department of Bioengineering have devised a method for repeatedly encoding, storing and erasing digital data within the DNA of living cells.
Sometimes, remembering and forgetting are hard to do.
"It took us three years and 750 tries to make it work, but we finally did it," said Jerome Bonnet, PhD, of his latest research, a method for repeatedly encoding, storing and erasing digital data within the DNA of living cells.
Bonnet, a postdoctoral scholar at Stanford University, worked with graduate student Pakpoom Subsoontorn and assistant professor Drew Endy, PhD, to reapply natural enzymes adapted from bacteria to flip specific sequences of DNA back and forth at will. All three scientists work in the Department of Bioengineering, a joint effort of the School of Engineering and the School of Medicine.
In practical terms, they have devised the genetic equivalent of a binary digit — a "bit" in data parlance. "Essentially, if the DNA section points in one direction, it's a zero. If it points the other way, it's a one," Subsoontorn explained.
"Programmable data storage within the DNA of living cells would seem an incredibly powerful tool for studying cancer, aging, organismal development and even the natural environment," said Endy.
Researchers could count how many times a cell divides, for instance, and that might someday give scientists the ability to turn off cells before they turn cancerous.
In the computer world, their work would form the basis of what is known as non-volatile memory — data storage that can retain information without consuming power. In biotechnology, it is known by a slightly more technical term, recombinase-mediated DNA inversion, after the enzymatic processes used to cut, flip and recombine DNA within the cell.
The team calls its device a "recombinase addressable data" module, or RAD for short. They used RAD to modify a particular section of DNA within microbes that determines how the one-celled organisms will fluoresce under ultraviolet light. The microbes glow red or green depending upon the orientation of the section of DNA. Using RAD, the engineers can flip the section back and forth at will.
They report their findings in a paper that will be published online May 21 in the Proceedings of the National Academy of Sciences. Bonnet is the first author of the paper, and Endy is the senior author.
To make their system work, the team had to control the precise dynamics of two opposing proteins, integrase and excisionase, within the microbes. "Previous work had shown how to flip the genetic sequence — albeit irreversibly — in one direction through the expression of a single enzyme," Bonnet said, "but we needed to reliably flip the sequence back and forth, over and over, in order to create a fully reusable binary data register, so we needed something different."
"The problem is that the proteins do their own thing. If both are active at the same time, or concentrated in the wrong amounts, you get a mess and the individual cells produce random results," Subsoontorn continued.
The researchers found it was fairly easy to flip a section of DNA in either direction. "But we discovered time and again that most of our designs failed when the two proteins were used together within the same cell," said Endy. "Ergo: Three years and 750 tries to get the balance of protein levels right."
Bonnet has now tested RAD modules in single microbes that have doubled more than 100 times and the switch has held. He has likewise switched the latch and watched a cell double 90 times, and set it back. The latch will even store information when the enzymes are not present. In short, RAD works. It is reliable and it is rewritable.
For Endy and the team, the future of computing then becomes not only how fast or how much can be computed, but when and where computations occur and how those computations might impact our understanding of and interaction with life.
"One of the coolest places for computing," Endy said, "is within biological systems."
His goal is to go from the single bit he has now to eight bits — or a "byte" — of programmable genetic data storage.
"I'm not even really concerned with the ways genetic data storage might be useful down the road, only in creating scalable and reliable biological bits as soon as possible. Then we'll put them in the hands of other scientists to show the world how they might be used," Endy said.
To get there, however, science will need many new tools for engineering biology, he added, but it will not be easy. "Such systems will likely be 10 to 50 times more complicated than current state-of-the-art genetic engineering projects," he said.
For what it is worth, Endy anticipates their second bit of rewritable DNA data will arrive faster than the first and the third faster still, but it will take time.
"We're probably looking at a decade from when we started to get to a full byte," he said. "But, by focusing today on tools that improve the engineering cycle at the heart of biotechnology, we'll help make all future engineering of biology easier, and that will lead us to much more interesting places."
More information: “Rewritable digital data storage in live cells via engineered control of recombination directionality,” Bonnet, J., Subsoontorn, P. & Endy, D. PNAS, http://dx.doi.org/ … s.1202344109 (2012).
Scientists from Stanford's Department of Bioengineering have devised a method for repeatedly encoding, storing and erasing digital data within the DNA of living cells.
Sometimes, remembering and forgetting are hard to do.
"It took us three years and 750 tries to make it work, but we finally did it," said Jerome Bonnet, PhD, of his latest research, a method for repeatedly encoding, storing and erasing digital data within the DNA of living cells.
Bonnet, a postdoctoral scholar at Stanford University, worked with graduate student Pakpoom Subsoontorn and assistant professor Drew Endy, PhD, to reapply natural enzymes adapted from bacteria to flip specific sequences of DNA back and forth at will. All three scientists work in the Department of Bioengineering, a joint effort of the School of Engineering and the School of Medicine.
In practical terms, they have devised the genetic equivalent of a binary digit — a "bit" in data parlance. "Essentially, if the DNA section points in one direction, it's a zero. If it points the other way, it's a one," Subsoontorn explained.
"Programmable data storage within the DNA of living cells would seem an incredibly powerful tool for studying cancer, aging, organismal development and even the natural environment," said Endy.
Researchers could count how many times a cell divides, for instance, and that might someday give scientists the ability to turn off cells before they turn cancerous.
In the computer world, their work would form the basis of what is known as non-volatile memory — data storage that can retain information without consuming power. In biotechnology, it is known by a slightly more technical term, recombinase-mediated DNA inversion, after the enzymatic processes used to cut, flip and recombine DNA within the cell.
The team calls its device a "recombinase addressable data" module, or RAD for short. They used RAD to modify a particular section of DNA within microbes that determines how the one-celled organisms will fluoresce under ultraviolet light. The microbes glow red or green depending upon the orientation of the section of DNA. Using RAD, the engineers can flip the section back and forth at will.
They report their findings in a paper that will be published online May 21 in the Proceedings of the National Academy of Sciences. Bonnet is the first author of the paper, and Endy is the senior author.
To make their system work, the team had to control the precise dynamics of two opposing proteins, integrase and excisionase, within the microbes. "Previous work had shown how to flip the genetic sequence — albeit irreversibly — in one direction through the expression of a single enzyme," Bonnet said, "but we needed to reliably flip the sequence back and forth, over and over, in order to create a fully reusable binary data register, so we needed something different."
"The problem is that the proteins do their own thing. If both are active at the same time, or concentrated in the wrong amounts, you get a mess and the individual cells produce random results," Subsoontorn continued.
The researchers found it was fairly easy to flip a section of DNA in either direction. "But we discovered time and again that most of our designs failed when the two proteins were used together within the same cell," said Endy. "Ergo: Three years and 750 tries to get the balance of protein levels right."
Bonnet has now tested RAD modules in single microbes that have doubled more than 100 times and the switch has held. He has likewise switched the latch and watched a cell double 90 times, and set it back. The latch will even store information when the enzymes are not present. In short, RAD works. It is reliable and it is rewritable.
For Endy and the team, the future of computing then becomes not only how fast or how much can be computed, but when and where computations occur and how those computations might impact our understanding of and interaction with life.
"One of the coolest places for computing," Endy said, "is within biological systems."
His goal is to go from the single bit he has now to eight bits — or a "byte" — of programmable genetic data storage.
"I'm not even really concerned with the ways genetic data storage might be useful down the road, only in creating scalable and reliable biological bits as soon as possible. Then we'll put them in the hands of other scientists to show the world how they might be used," Endy said.
To get there, however, science will need many new tools for engineering biology, he added, but it will not be easy. "Such systems will likely be 10 to 50 times more complicated than current state-of-the-art genetic engineering projects," he said.
For what it is worth, Endy anticipates their second bit of rewritable DNA data will arrive faster than the first and the third faster still, but it will take time.
"We're probably looking at a decade from when we started to get to a full byte," he said. "But, by focusing today on tools that improve the engineering cycle at the heart of biotechnology, we'll help make all future engineering of biology easier, and that will lead us to much more interesting places."
More information: “Rewritable digital data storage in live cells via engineered control of recombination directionality,” Bonnet, J., Subsoontorn, P. & Endy, D. PNAS, http://dx.doi.org/ … s.1202344109 (2012).
Sol Luckman graphic
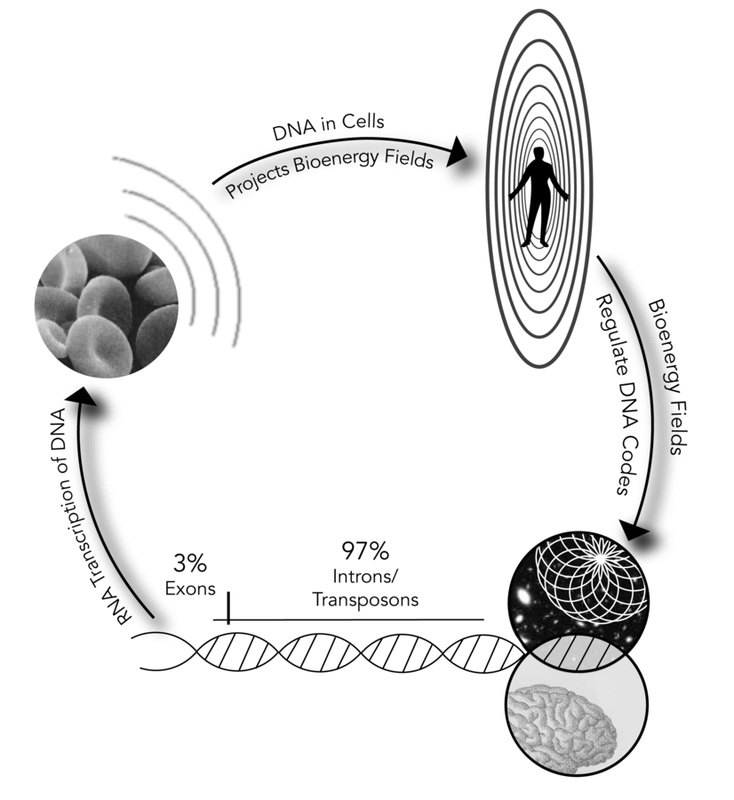
The above diagram illustrates how body building is both genetic, involving RNA transcription of DNA codes to create cells, and energetic, dependent on the interface between the bioenergy fields and “junk” or potential DNA for regulation of cellular composition. This diagram also shows how potential DNA’s transposons can be prompted directly by consciousness, internal (personal) and external (universal), to modify cellular replication.
(Image and text from CONSCIOUS HEALING: BOOK ONE ON THE REGENETICS METHOD)
The above diagram illustrates how body building is both genetic, involving RNA transcription of DNA codes to create cells, and energetic, dependent on the interface between the bioenergy fields and “junk” or potential DNA for regulation of cellular composition. This diagram also shows how potential DNA’s transposons can be prompted directly by consciousness, internal (personal) and external (universal), to modify cellular replication.
(Image and text from CONSCIOUS HEALING: BOOK ONE ON THE REGENETICS METHOD)
Epigenetics:
The Dogma-defying Discovery That Genes Learn From Experience
Haley Peckham.
MA (Philosophy), MSc (Neuroscience) doi: 10.12744/ijnpt.2013.0009-0020
Introduction
Epigenetics challenges our acceptance of the dichotomy between nature and nurture. It is coming to be understood that nurture somehow writes its way into our genes; that experiences, or the effects of experiences on gene transcription, may continue throughout our own lives and into those of the next generation. For anyone engaged in the challenge of helping clients who have suffered traumatic, neglectful or abusive experiences, or for anyone offering clients a new experience through the psychotherapeutic relationship, an understanding of epigenetics is pertinent and illuminating. Here I introduce the concept of epigenetics, describe some epigenetic mechanisms, and review the findings of a number of recent studies that seek to understand why, what, and how our genes can learn from experience. The engaging debate of whether it is nature or nurture that has the most influence over who we are and what kind of lives we lead often illuminates the attitudes the debaters hold with regard to their own lives. In my early twenties, I tended to champion the influence of the environment, as although I felt affected by childhood adversity, I did not like to think of myself as passively, fatalistically accepting nature—my genetic inheritance—as my lot in life. Discoveries in biology have made this debate and my wrangling with it entirely passé with the enthralling field of epigenetics, which allows for degrees of gene expression or “shades of grey” that release us from the “black and white” dichotomy of genetic determinism. Epigenetics provides insights into how the environment dynamically impacts on our gene expression as humans, influencing, amongst other traits, our and our children’s health, ability to learn and remember, and responses to stress. The fabric of the lives led by our parents and grandparents, from their diet (Katada, Imhof, & Sassone-Corsi, 2012) to their education, the care they received, traumas they may have suffered, and perhaps many other experiences leave their legacy written alongside our DNA as “instructions for interpretation.” The purpose of this as yet inappreciably sophisticated and elegant code of epigenetic information, if it can be properly be called a “purpose,” is to record, use, and pass on information about the intricacies of the environment experienced in both our lives and those that have gone before us, so that we and our children can be as well prepared for survival in our particular environment as it possible to be. Life, over lives, continually perceives the environment, and the epigenetic instructions for interpreting the DNA are written, used, revised, and may be inherited, along with our genes. Epigenetics is the contemporary study of how the environment influences gene expression both within and, through heritable changes in DNA, beyond the lifetime of an organism. The idea that organisms can inherit environmentally acquired characteristics is, however, the old idea of Lamarckian inheritance. In 1809, Jean-Baptiste Lamarck suggested that an organism would acquire traits through adapting to its environment, and that these traits would then be inherited by its offspring (Handel & Ramagopalan, 2010). Lamarck’s theory was overlooked in favor of Darwin’s natural selection theory of evolution, as the two explanations appeared at the time to be mutually exclusive, but the advent of epigenetics has made it possible for these theories to be reconciled.
“Epigenetics” literally means “above the genes,” and is the means by which the environment “marks” the genes, dramatically or subtly, changing their level of expression either transiently or for our lifetime, or, through inheritability, throughout our children’s and grandchildren’s lifetimes. A formal definition of epigenetic events proposed by Adrian Bird is: “the structural adaption of chromosomal regions so as to register, signal or perpetuate altered activity states” (Bird, 2007, p. 398). This definition encompasses the broad remit of epigenetic marks from transient, where the epigenetic mark ascribed by the environmental adaption lasts only a few hours, to heritable, where the environmental effects last over a generation. The brain-derived neurotrophic factor (BDNF) gene, implicated in psychiatric disorders and learning and memory (Autry & Monteggia, 2012; Boulle et al., 2012; Lu, Christian, & Lu, 2008; Minichiello, 2009; Musumeci & Minichiello, 2011), is subject to both short- and long-term epigenetic marking in rodents. For example, following the favorable social experience of being reared in a communal nest, mice challenged with one hour in a mildly stressful novel environment generate hippocampal BDNF faster than mice raised in a standard nest, as a result of an epigenetic mark on the BDNF gene (Branchi, Karpova, D’Andrea, Castren, & Alleva, 2011). However, rat pups subjected to a rat equivalent of childhood maltreatment are epigenetically marked by this experience, reducing the level of BDNF in their pre-frontal cortex throughout their adult life. The offspring of these rats also carry the same epigenetic mark on their BDNF gene even when they have been cross-fostered to non-maltreating mothers (Roth, Lubin, Funk, & Sweatt, 2009). Thus, epigenetics, the mechanisms by which our genes record or adapt to the environment, can shape gene expression over a few minutes, an hour, or a lifetime, and can even shape the gene expression pattern of the next generation. It is even possible for genes to “remember” an event and make a contingency plan for its recurrence, as in the case of the corticotropin-releasing hormone gene of rat pups. In response to maternal deprivation, the promoter region of this gene is epigenetically marked. Later, following a stressful experience in adulthood, the pre-recorded epigenetic mark leads to a hypersensitive stress response, observed as a more actively transcribed corticotropin-releasing hormone gene and increased levels of the stress hormone corticosterone (Chen et al., 2012). There are three main types of epigenetic modification: DNA methylation and histone modification, outlined here together with examples of these mechanisms in action, and translational regulation by micro RNA (not pertinent to this review, but see (Haramati et al., 2011; Sato, Tsuchiya, Meltzer, & Shimizu, 2011) for useful introductions). Types of epigenetic modification, Level I: DNA methylation Methylation involves the addition of a methyl group to mammalian DNA, and can occur in response to environmental influence, making a stable, potentially heritable addition to the DNA that enhances or represses the transcription of a gene. Methylation is not a mutation, as the sequence of bases (adenine, guanine, cytosine, and thymine) remains the same. The methyl group is added to a cytosine nucleotide, usually followed by a guanine nucleotide (a CpG site), by enzymes known as DNA methyltransferases (DNMTs). The promoter regions or transcriptional start site of genes (the point at which RNA polymerase II begins transcription of the DNA into mRNA) are frequently rich in CpG sites, and methylation of these regions is associated with long-term silencing of genes. Methylation of CpG sites within the body of the gene is more ambiguous and context specific and can lead to either repression or activation (Jones, 2012)’. Methylated DNA can also be bound by Methyl Binding Domain (MBD) proteins such as MeCP2 which recruit other protein complexes to re-model the local chromatin (DNA + histone packaging), leading either to repression or activation of specific genes. MeCP2 is an important point of integration between types of epigenetic modification, as MeCP2 binds methylated DNA but also can recruit chromatin re-modeling proteins which implement the other major type of epigenetic alteration, histone modification, described below (Cedar & Bergman, 2012). Methylation of DNA is reversible but instead of de-methylation occurring via the action of a single enzyme, it occurs through a multistep process. It is intriguing that almost half of the methylated DNA in the brain is 5-hydroxymethylcytosine, an intermediate formed during the multi-step de-methylation of DNA (Szulwach et al., 2011), a process which is dependent on neuronal activity (Guo, Su, Zhong, Ming, & Song, 2011). The contemporary view of DNA methylation is that, in the brain especially, it is a highly dynamic process, responsive to neural activity (and therefore experience) and crucial for plasticity-related functions such as learning, memory and behavioral adaptation (Baker-Andresen, Ratnu, & Bredy, 2013). Types of epigenetic modification, Level II: Histone modification The DNA in the nucleus of each eukaryotic cell, including in the case of humans, contains all of the genetic material. So a human skin cell carries the same complete genetic code as a neuron or a parietal cell of the stomach. Each specialized cell need only be able to transcribe a small fraction of the total DNA it carries in order to perform its specific function. Since the rest of its DNA will never need to be accessed, it can be packed away as efficiently as possible. Histone proteins facilitate this efficient packaging of DNA, largely due to the electrostatic attraction between the negatively charged DNA and the positively charged histone proteins. DNA is coiled around a core octamer of histone proteins, like thread on a spool, forming a single nucleosome. Multiple nucleosomes together are known as chromatin, which has different conformations reflecting the accessibility of the genes for transcription. Euchromatin is open chromatin, conceivable as separate beads on a string; here the genes are accessible, whereas in condensed chromatin or heterochromatin, the genes are inaccessible and un-transcribed, as they are so densely packed (Jenuwein & Allis, 2001).
Figure 1. Structure of a nucleosome: DNA wound around a histone core. From “Epigenetic Regulation in Psychiatric Disorders,” by N. Tsankova, W. Renthal, A. Kumar, and E. J. Nestler, 2007, Nature Reviews Neuroscience, 8, p. 356. Copyright 2007 by Nature Publishing Group. Reproduced with permission. Like most proteins, the histone proteins comprising the core around which the DNA winds itself can have chemical groups attached to them that may either subtly change their charge, and thus how tightly they are bound to the DNA, or change their affinity for other protein-binding partners. In this way, histone modifications alter the chromatin structure and hence the accessibility of genes, but can also affect the binding of other molecules including DNA methyltransferases—the enzymes that add methyl groups to DNA, not the histone proteins that the DNA coils around (Ooi et al., 2007). This interaction forms another point of integration between DNA-modifying and histone-modifying epigenetic mechanisms. Histones may be acetylated, methylated (including bi- or tri-methylated), phosphorylated or ubiquitinylated on specific residues, and each group or combination of groups may signal a precise outcome. For example, an epigenetic modification of a tri-methylated lysine4 residue on histone3 signals an actively transcribed gene, whereas a tri-methylation on lysine27 of histone3 signals a repressed gene state. If lysines 4 and 27 are both tri-methylated, the gene is thought to be “poised,” ready for activation (Bernstein et al., 2006). As there are four histones (two of each make up the octamer core), and at least four chemical groups that can attach to multiple residues, there are a large number of possible combinations of epigenetic marks that can be made on a histone octamer, giving rise to the concept of a “histone code” that can nuance gene expression (Jenuwein & Allis, 2001). There are enzymes that catalyze the addition and removal of each type of chemical group. Histone acetylases (HATs), for example, add acetyl groups to the histone, reducing the electrostatic charge difference and therefore the affinity between the histone and the DNA, thus loosening the coil. Accordingly, HATs tend to be associated with gene activation, whereas histone de-acetylases (HDACs) remove the acetyl group, increase the electrostatic attraction, tighten the coil, and are thus considered transcriptional repressors (Bannister & Kouzarides, 2011). In contrast to the epigenetic modification of a methylation made to DNA, histone modifications are easily reversible and highly dynamic, although the enzymes that catalyze the addition and removal of various groups are themselves regulated. The histone code also has great flexibility in determining how long a gene’s activity may be enhanced or repressed, as epigenetic marks may record an experience on a gene without necessarily changing its level of transcription. They may transiently enhance or repress gene expression, or may recruit (or repel) DNA methylases that can administer de novo methylations on the DNA which result in the gene becoming stably repressed throughout the life of the organism and in the next generation (Ooi, et al., 2007). It is interesting to note that Valproic acid, a known teratogen used in the treatment of bipolar disorder, is a histone de-acetylase inhibitor but also leads to reduced levels of DNA methylation, demonstrating that a transient and easily reversible epigenetic histone acetylation is frequently followed by a more stable and less easily reversed DNA methylation, silencing the gene (Alonso-Aperte, Ubeda, Achon, Perez-Miguelsanz, & Varela-Moreiras, 1999; Gottlicher, 2004). Thus, histone modification and DNA methylation, whilst distinct epigenetic mechanisms, can act in synergy to produce permanent gene silencing. Figure 2. The dynamic addition and removal of chemical groups to histone tails by specific enzymes. From “Epigenetic Regulation in Psychiatric Disorders,” by N. Tsankova, W. Renthal, A. Kumar, and E. J. Nestler, 2007, Nature Reviews Neuroscience, 8, p. 356. Copyright 2007 by Nature Publishing Group. Reproduced with permission. The epigenetic impact of early maternal care Many animal studies of epigenetics have looked at the impact of early maternal care on the hypothalamic-pituitary-adrenal (HPA) axis or, put more simply, the stress axis. This axis co-ordinates the response to stress from the brain, which perceives the stress, to the adrenal gland, which releases glucocorticoids such as cortisol (corticosterone in rodents) in response to the stressor. The hypothalamus receives neural input—our perception of the stress—and releases corticotropin-releasing hormone (CRH) that acts on the anterior pituitary gland to release adrenocorticotropic hormone (ACTH). This in turn acts on the cortex of the adrenal gland to release cortisol. The multiple steps in this process allow multiple opportunities for the glucocorticoids to negatively feed back at several levels of the HPA axis, including the hippocampus and the hypothalamus, limiting CRH release, and at the pituitary, limiting ACTH release (Anacker, Zunszain, Carvalho, & Pariante, 2011; Pariante & Lightman, 2008). Glucocorticoids feed back via glucocorticoid receptors. More receptors means a swifter, more efficient negative feedback mechanism resulting in lower levels of circulating CRH, ACTH and cortisol, and consequently an HPA axis that readily returns to homeostasis. However, in response to chronically high levels of circulating glucocorticoids, glucocorticoid receptors are down-regulated; as a result, the negative feedback mechanism is less efficient, and the HPA axis is slower to return to homeostasis or may become dysregulated (Anacker, et al., 2011; Sapolsky, Meaney, & McEwen, 1985). The HPA axis is an adaptive system subject to early-life epigenetic programming both prenatally and postnatally through variations in maternal care, separation and abuse. Early pre-natal stress in mice has been epigenetically linked to enduring changes in HPA axis reactivity and a depressive phenotype in male offspring. Male adult mice whose mothers had been mildly stressed early in utero had higher levels of CRH and fewer glucocorticoid receptors in their hippocampus, the former increasing the activation of the HPA axis and the latter reducing the ability to regulate the HPA axis. These mice exhibited behaviors associated with depression and, following exposure to restraint stress, had increased levels of corticosterone compared to mice whose mothers had not been subjected to early pre-natal stress. When the promoter regions of relevant genes were examined, it was discovered that the promoter for corticotropin-releasing hormone was hypo-methylated, increasing the expression of CRH, the hormonal activator of the HPA axis. The glucocorticoid receptor promoter was also hyper-methylated, reducing its level of expression and thereby the mouse’s ability to negatively regulate its HPA axis after activation by CRF. This would at least contribute to, if not cause, the observed depressed behavior (Mueller & Bale, 2008). Figure 3. Combinations of histone marks corresponding to active, permissive, repressed or inactive (silenced) genes. From “Epigenetic Regulation in Psychiatric Disorders,” by N. Tsankova, W. Renthal, A. Kumar, and E. J. Nestler, 2007, Nature Reviews Neuroscience, 8, p. 356. Copyright 2007 by Nature Publishing Group. Reproduced with permission. Early post-natal stress has also been linked to enduring epigenetic changes that alter the reactivity of the HPA axis. Rat pups that receive enhanced licking and grooming by their mothers in the first ten days of life acquire a permanent increase in their number of glucocorticoid receptors, meaning that their HPA axis is more readily negatively regulated. Thus, when these pups become adults and are exposed to an acute stressor, they release corticosterone, which negatively feeds back through a greater number of glucocorticoid receptors, quickly reducing the levels of CRH, circulating ACTH, and corticosterone (Liu et al., 1997). If this effect was correlated with measurable changes to the epigenome, i.e., DNA methylation or histone code changes at the site of a relevant gene, it would support the hypothesis that maternal care leads to epigenetic changes that moderate the activity of the stress axis. In support of this hypothesis, a landmark study by Weaver’s group (Weaver, Diorio, Seckl, Szyf, & Meaney, 2004) compared rat pups that had either high or low levels of licking and grooming from mothers with whom they were cross-fostered, and found epigenetic differences in the promoter region of the glucocorticoid receptor gene. Pups that had received high levels of licking and grooming had hypo-methylated DNA and hyper-acetylated histone3 at the promoter region of the glucocorticoid receptor gene where a transcription factor binds, leading to enhanced transcriptional activity of the glucocorticoid receptor gene and therefore to more glucocorticoid receptors. As adults, pups that had received low levels of licking and grooming had fewer glucocorticoid receptors and correspondingly higher levels of circulating corticosterone following restraint stress. Taking the study a step further towards identifying maternal care as a cause of epigenetic changes that modify the reactivity of the HPA axis, a histone de-acetylase (HDAC) inhibitor was administered to the rats which not only enhanced acetylation of the glucocorticoid promoter in the low licking and grooming group, it also removed all of the other observed differences in this group: the glucocorticoid promoter became hypo-methylated, more glucocorticoid receptors were expressed, and the corticosterone response following restraint stress was normalized to the same level as the high licking and grooming group. Essentially, the epigenetic differences brought about by lower levels of maternal care were reversed by the HDAC inhibitor (Weaver et al., 2004). In a follow-up study by the same group (Weaver et al., 2005), rats were infused with a methyl group donator. Even if the rats had received high levels of licking and grooming by their mothers when they were pups, leading to hypo-methylation of the glucocorticoid promoters, if they received the methylating drug as adults, all of the beneficial effects of this extra licking and grooming were reversed; the glucocorticoid promoter became hyper-methylated and fewer glucocorticoid receptors were expressed, leading to higher levels of corticosterone following a stressful experience and higher scores in a test of behavioral responses to stress (Weaver et al., 2005). From their combined studies, the authors primarily concluded that maternal behavior programs the stress response through epigenetic modification of the glucocorticoid promoter, and secondarily concluded that although maternal care makes stable changes to the epigenome, these are pharmacologically modifiable even in adults (Weaver, et al., 2005). Since then, in the case of studies showing that differences in estrogen receptor levels correlated with the level of maternal care received as a pup and the level of maternal care given in turn to offspring (Champagne, Weaver, Diorio, Sharma, & Meaney, 2003), the relationship has also been found to be mediated through epigenetic changes to the estrogen receptor promoter. Thus, receiving a high level of maternal care leads to reduced methylation of the CpG region within the promoter of the estrogen receptor, correspondingly increased levels of expressed estrogen receptors, and the capacity to give a high level of maternal care to the following generation of pups (Champagne et al., 2006). Separation from mother in infancy has long been known to have emotionally and psychologically detrimental sequelae, and the work of two brilliant and influential researchers, Harry Harlow and John Bowlby, is eloquently discussed in van der Horst and van der Veer’s review (van der Horst & van der Veer, 2008). In the field of epigenetics, the effects of maternal separation on the reactivity of the HPA axis are beginning to be discovered. In the HPA axis, stressful stimuli perceived at the level of the hypothalamus lead to secretion of CRH-stimulating ACTH. Arginine vasopressin (AVP) is also secreted by the hypothalamus, and potentiates the effect of CRH at the anterior pituitary, stimulating more ACTH release and increased HPA activity. Mice that were separated from their mothers for three hours a day for the first ten days of their lives had higher basal levels of corticosterone as well as higher levels following a stressful experience, and also scored higher on behavioral measures of stress and lower on memory tasks. Higher levels of expressed AVP correlated with hypo-methylated DNA in the regulatory region of the AVP gene (Murgatroyd et al., 2009). The methyl-binding protein MeCP2, which binds methylated DNA, ordinarily represses AVP transcription. But in mice that have been separated from their mothers, MeCP2 cannot bind as readily to the AVP promoter, as it is hypo-methylated even when the mouse reaches adulthood, leading to reduced repression by MeCP2 and consequent higher levels of AVP transcription. This results in a persistent hormonal activation of the HPA axis throughout the life of the mouse (Murgatroyd, et al., 2009). Figure 4. The hypothalamic pituitary adrenal axis and its regulation by glucocorticoid negative feedback. Illustration by Brian M. Sweis (2012). Reproduced from http://commons.wikimedia.org/wiki/File:HPA_Axis_Diagram_(Brian_M_Sweis_2012).png As well as enduringly activating the HPA axis, early life stress induced by maternal separation has been shown to enhance performance in stress-related memory tasks in young rats (Suri et al., 2012). The rats’ improved performance correlated with increased neurogenesis, reduced repressive histone methylation of a BDNF gene promoter, and correspondingly higher levels of BDNF in the hippocampus, compared to controls. Interestingly, once the maternally separated rats reached middle age, all these beneficial effects were lost, although this could be ameliorated by long-term antidepressant treatment. The authors of this study suggested that maternal separation stress induces biologically adaptive responses including epigenetic changes that increase the rat pups’ chances of having to survive alone, without the mother’s help and protection, but that these adaptive changes exact a heavy toll when the rat reaches middle age (Suri, et al., 2012). From an evolutionary point of view, as long as the adaptive response to maternal separation allows the rat to survive on its own long enough to be able to reproduce, the exacted toll is worthwhile. Paradigms that mimic abusive behavior in humans have also been used to examine the epigenomic effects of maternal abuse in rodents, demonstrating that abuse leads to significant hypo-methylation of the BDNF promoter in the pre-frontal cortex. Notably, rats that had suffered maltreatment in infancy also maltreated their own (or cross-fostered) pups, and the effect of their own maltreatment could be seen in their biological offspring as a hypo-methylated BDNF promoter region even when the pups were fostered to a caring mother (Roth, et al., 2009). Maltreatment, or abuse, appears to be such a profound environmental influence that its effects linger in the generation following that which experienced the maltreatment firsthand. A similar observation of the trans-generational transmittance of a profound experience has been made by Rachel Yehuda, a researcher into post-traumatic stress disorder (PTSD) who found a higher than normal prevalence of PTSD in the children of mothers who had PTSD as a result of the Holocaust, suggesting a mechanism other than traditional genetic contribution in play, with epigenetics being the obvious candidate (Yehuda, Bell, Bierer, & Schmeidler, 2008). Although specific epigenetic modifications have not yet been linked to experiences that lead to the development of PTSD in humans, a recent study has identified patterns of genes that are differentially methylated in human sufferers of PTSD (Uddin et al., 2010). Additionally, in the rat model of PTSD, where the animals are exposed on two occasions to a cat, as well as having a new rat cage mate daily for 31 days, dramatic epigenetic changes have been observed in the promoter region of the BDNF gene in the hippocampus (Roth, Zoladz, Sweatt, & Diamond, 2011). Taken together, these results suggest that the development of human PTSD may occur through traumatic events causing epigenetic changes to specific genes, which, as well as leading to PTSD in the trauma victim, could confer an increased risk of the same in the victim’s children, as the epigenetic signatures on the genes caused by the trauma could be inherited (Yehuda & Bierer, 2009). Although speculative, it seems likely that the inheritance of genes that have been epigenetically marked by trauma could be biologically adaptive and enhance the chance of survival if the environment remains life-threatening for the next generation. Hyper-vigilance and extreme alertness following trauma, whilst profoundly exhausting and distressing, may be life-saving in a dangerous environment. Human studies looking at the epigenetic impact of the environment Gathering data for human studies tends to involve the use of postmortem tissue and detailed retrospective analyses of significant life events or trauma. A recent study using these methodologies compared the differences in DNA methylation across the whole genome of 25 men with a history of severe child abuse compared to 16 controls, and found that fewer genes are actively transcribed in men who suffered abusive childhoods. The methylation profiles of the men showed 248 genes were hyper-methylated and 114 genes were hypo-methylated in the group with a history of abuse compared to the control group (Labonte et al., 2012). Of these, the top five most hyper-methylated genes were neuronal, and many of the genes that were differently methylated in the group who had experienced childhood abuse were related to plasticity, i.e., genes that are known to have a role in learning or adaptive mechanisms. A similar study using blood samples found that children raised in institutions had more methylated, and thus fewer expressed, genes than children raised by their biological parents (Naumova et al., 2012). It is tempting to speculate that perhaps lower levels of care restrict the variety of genes that can be expressed, whereas higher levels of care are the epigenetic gateway to our available genome. We speak of wanting to give our children “every opportunity”, but the reality of what epigenetic opportunities are available to our children could largely depend on the care we give them very early in life. Access to the widest potential of their genome could be profoundly influenced by their experience of our care, and if the genes that are accessed by enhanced care perform plasticity-related functions, the effect of that care is compounded for good or ill. Poor care may mean that fewer plasticity-related genes are expressed, thus restricting a child (and the adult’s) potential to learn, remember or adapt within their environment, manifesting in less flexibility and perhaps more stereotyped or rigid responses. Studies that identify the epigenetic effects of levels of care or experiences on specific genes have been undertaken in humans. In a striking human parallel to Weaver’s (2004) rodent study showing that low levels of maternal care epigenetically modified the glucocorticoid receptor promoter by hyper-methylation, reducing the transcription of the glucocorticoid receptor gene, McGowan et al. (2009) found that suicide victims who had a history of childhood abuse had hyper-methylated glucocorticoid promoters and fewer HPA axis-regulating hippocampal glucocorticoid receptors than controls. Both prenatal stress and even the birth experience have also been linked to epigenetic changes. The depressive mood of mothers in the third trimester of pregnancy has been linked to hyper-methylated glucocorticoid receptor promoters and higher cortisol responses in three-month-old babies (Oberlander et al., 2008), while infants born by Caesarean section have significantly higher levels of DNA methylation in their leucocytes compared to vaginal births at the time of delivery, although this normalizes after 3–5 days (Schlinzig, Johansson, Gunnar, Ekstrom, & Norman, 2009).
Pharmacotherapeutics, enriching environments, and reversible epigenetic change
Pharmacological treatments can reverse epigenetic marks caused by environmental experiences on both the BDNF gene (Roth, et al., 2009; Suri, et al., 2012) and the glucocorticoid receptor gene (Weaver, et al., 2005; Weaver, Meaney, & Szyf, 2006). Antidepressant treatment has frequently been shown to restore levels of BDNF in animal models of depression (Balaratnasingam & Janca, 2012), and in the social defeat, mouse model of depression, imipramine has been demonstrated to act through epigenetic mechanisms, reversing the down-regulating effect that social defeat has on BDNF transcription levels, although it does so by a compensatory HDAC-inhibiting mechanism rather than direct reversal of repressive histone methylations (N. M. Tsankova et al., 2006). The HDAC inhibitor, valproic acid, a treatment in certain psychiatric disorders, also epigenetically enhances BDNF transcription, facilitating the forgetting (or extinction) of fear-conditioned learning (Bredy et al., 2007; Whittle et al., 2013). Other histone-modifying agents may become useful psychoactive medications. It has recently been demonstrated that the acetylating agent L-acetyl-carnitine, already available as a dietary supplement, enhances transcription of the type II metabotropic glutamate receptor, which has a swift and enduring anti-depressant effect in rodents (Nasca et al., 2013). As epigenetic mechanisms mediate the effects of environment, the involvement of epigenetic mechanisms in psychiatric disorders that have an environmental component is axiomatic. Whilst pharmacotherapeutics holds promise for the treatment of psychiatric disorders (Grayson, Kundakovic, & Sharma, 2010; N. Tsankova, Renthal, Kumar, & Nestler, 2007), for the many people who fall below the threshold of a psychiatric diagnosis or who prefer non-medicating treatments, the very nature of the epigenetic mechanism suggests an alternative to drug treatment in the form of new, desirable environmental experiences to overwrite previous negative or undesirable experiences written on the epigenome. Environmental enrichment is proof of the principle that new environmental experiences can reverse or ameliorate the epigenetic effects of a previously impoverished or stressful environment. Enriching environments provide opportunities for enhanced sensory, motor, cognitive, and social stimulation, and have a multitude of beneficial effects, from enhancing neural plasticity and learning and memory, to conferring resilience to depression and ameliorating the effects of many brain disorders including Alzheimer’s and Huntington’s diseases and stroke (Bekinschtein, Oomen, Saksida, & Bussey, 2011; D’Andrea, Gracci, Alleva, & Branchi, 2010; Nithianantharajah & Hannan, 2006). The effects of an enriched environment in rodents are profound, and can even mitigate the effects of prenatal stress or maternal separation on the HPA axis (Francis, Diorio, Plotsky, & Meaney, 2002; Morley-Fletcher, Rea, Maccari, & Laviola, 2003). Analogously to the effects of trauma, the effects of enrichment can be seen in the next generation, improving learning and memory (Arai, Li, Hartley, & Feig, 2009) and, following the socially enriching experience of being reared in a communal nest, enhancing levels of maternal care in the next generation, reducing anxiety behavior, and increasing litter sizes (Curley, Davidson, Bateson, & Champagne, 2009). It is likely that epigenetic mechanisms mediate the benefits of enrichment in terms of both the generation experiencing the enrichment and the trans-generational effects of enrichment that are observed. Recent work shows that rats whose parents had enriching experiences have less methylated DNA than rats whose parents experienced standard housing (Mychasiuk et al., 2012), suggesting that enrichment (such as high levels of care) increases the number of genes that will be actively transcribed. This perhaps reflects a need for greater biological complexity in order to be able to thrive in a more complex (enriched) environment. One gene that is extremely responsive to the environment is BDNF, which is epigenetically modified by the experience of abuse (see above) as well as by the experience of an enriched environment. A recent study demonstrated increased levels of permissive histone methylations, enhancing transcription of the gene and raising levels of hippocampal BDNF in mice that experienced a month in an enriched environment (Kuzumaki et al., 2011). The role of BDNF in learning and memory behaviors, combined with its epigenetically regulated transcription, suggests this gene is a critical transducer of the experienced environment. Environmental enrichment is a powerful tool to effect epigenomic changes, which subsequently affect the levels of BDNF and modulate HPA axis activity, which, in turn, builds resilience to, or delays the onset of, various psychiatric or neurodegenerative disorders.
The relevance of epigenetics for
psychotherapy
When I learned that real-life subjective experiences could regulate gene transcription, it changed the way I understood myself. I was not just an immutable collection of transcribed genes. My genes had my experiences of early care, my traumas and dramas and education written onto them in the tiniest molecular inscription of the histone code and DNA methylations. Furthermore, even as I was born, the most salient facts of my parents’ and even their parents’ lives—the kinds of care they had received, their education, their sense of safety or fear—was etched into my genes too, to help me adapt to what was biologically anticipated to be a similar environment. I did not know about epigenetics when I underwent psychotherapy, but if I had, it may have inspired my curiosity about my parents’ lives and informed my developing narrative that who they were, who I was, was a more intricate and elegantly entwined interplay between our genes and our environments than I could ever have imagined. Neither they nor I was responsible or to blame; instead, we each were simply trying to adapt to our niche, trying to make the best of what we had: our genes and our unknown, influential, epigenetic legacy. In writing this article I wanted to share the concept of epigenetics and to review and communicate the most relevant findings in contemporary epigenetics research with psychotherapists or anyone who seeks to understand and alleviate emotional and psychological distress.
Epigenetics is an astounding and revolutionary discovery that brings compassionate understanding and great hope for the future to anyone who suffers distress, as well as those who work so intimately with distressed clients. An appreciation of epigenetics inspires compassion for those of us who come into the world vigilant and defensive, our epigenetic legacy anticipating a harsh world, a dangerous place to live; yet it also inspires hope, for even the most epigenetically determined aggressive, defensive stance is, by its very nature, responsive to the novelty of benign and benevolent environments.
Fundamentally, epigenetics translates experience in the world into a gene expression profile that shapes who we are. If we are to develop or change ourselves, we must actively seek the experiences that will help us develop along the trajectory we desire. Psychotherapy can be seen as a form of emotional, environmental enrichment. It is perfectly placed to offer an enriched experience of a caring relationship, albeit an asymmetrical one, where one can received empathic, attuned attention that may be wholly novel and life changing. By offering a new, benevolent experience of relationship, psychotherapy can be reasonably assumed to change gene expression, calming a hyper-reactive stress axis and ameliorating the effects of an epigenetically scarred BDNF promoter, and in so doing, changing our responses to stress and perhaps our capacity to learn and flexibly respond to emotionally challenging situations. For the client in psychotherapy, as in many situations in life, it really is the experience that counts.
References Alonso-Aperte, E., Ubeda, N., Achon, M., Perez-Miguelsanz, J., & Varela-Moreiras, G. (1999). Impaired methionine synthesis and hypomethylation in rats exposed to valproate during gestation. Neurology, 52(4), 750-756. Anacker, C., Zunszain, P. A., Carvalho, L. A., & Pariante, C. M. (2011). The glucocorticoid receptor: pivot of depression and of antidepressant treatment? Psychoneuroendocrinology, 36(3), 415-425. doi: 10.1016/j.psyneuen.2010.03.007 Arai, J. A., Li, S., Hartley, D. M., & Feig, L. A. (2009). Transgenerational rescue of a genetic defect in long-term potentiation and memory formation by juvenile enrichment. J Neurosci, 29(5), 1496-1502. doi: 10.1523/jneurosci.5057-08.2009 Autry, A. E., & Monteggia, L. M. (2012). Brain-derived neurotrophic factor and neuropsychiatric disorders. Pharmacol Rev, 64(2), 238-258. doi: 10.1124/pr.111.005108 Baker-Andresen, D., Ratnu, V. S., & Bredy, T. W. (2013). Dynamic DNA methylation: a prime candidate for genomic metaplasticity and behavioral adaptation. Trends Neurosci, 36(1), 3-13. doi: 10.1016/j.tins.2012.09.003 Balaratnasingam, S., & Janca, A. (2012). Brain Derived Neurotrophic Factor: a novel neurotrophin involved in psychiatric and neurological disorders. Pharmacol Ther, 134(1), 116-124. doi: 10.1016/j.pharmthera.2012.01.006 Bannister, A. J., & Kouzarides, T. (2011). Regulation of chromatin by histone modifications. Cell Res, 21(3), 381-395. doi: 10.1038/cr.2011.22 Bekinschtein, P., Oomen, C. A., Saksida, L. M., & Bussey, T. J. (2011). Effects of environmental enrichment and voluntary exercise on neurogenesis, learning and memory, and pattern separation: BDNF as a critical variable? Semin Cell Dev Biol, 22(5), 536-542. doi: 10.1016/j.semcdb.2011.07.002 Bernstein, B. E., Mikkelsen, T. S., Xie, X., Kamal, M., Huebert, D. J., Cuff, J., . . . Lander, E. S. (2006). A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell, 125(2), 315-326. doi: 10.1016/j.cell.2006.02.041 Bird, A. (2007). Perceptions of epigenetics. Nature, 447(7143), 396-398. doi: 10.1038/nature05913 Boulle, F., van den Hove, D. L., Jakob, S. B., Rutten, B. P., Hamon, M., van Os, J., . . . Kenis, G. (2012). Epigenetic regulation of the BDNF gene: implications for psychiatric disorders. Mol Psychiatry, 17(6), 584-596. doi: 10.1038/mp.2011.107 Branchi, I., Karpova, N. N., D’Andrea, I., Castren, E., & Alleva, E. (2011). Epigenetic modifications induced by early enrichment are associated with changes in timing of induction of BDNF expression. Neurosci Lett, 495(3), 168-172. doi: 10.1016/j.neulet.2011.03.038 Bredy, T. W., Wu, H., Crego, C., Zellhoefer, J., Sun, Y. E., & Barad, M. (2007). Histone modifications around individual BDNF gene promoters in prefrontal cortex are associated with extinction of conditioned fear. Learn Mem, 14(4), 268-276. doi: 10.1101/lm.500907 Cedar, H., & Bergman, Y. (2012). Programming of DNA methylation patterns. Annu Rev Biochem, 81, 97-117. doi: 10.1146/annurev-biochem-052610-091920 Champagne, F. A., Weaver, I. C., Diorio, J., Dymov, S., Szyf, M., & Meaney, M. J. (2006). Maternal care associated with methylation of the estrogen receptor-alpha1b promoter and estrogen receptor-alpha expression in the medial preoptic area of female offspring. Endocrinology, 147(6), 2909-2915. doi: 10.1210/en.2005-1119 Champagne, F. A., Weaver, I. C., Diorio, J., Sharma, S., & Meaney, M. J. (2003). Natural variations in maternal care are associated with estrogen receptor alpha expression and estrogen sensitivity in the medial preoptic area. Endocrinology, 144(11), 4720-4724. doi: 10.1210/en.2003-0564 Chen, J., Evans, A. N., Liu, Y., Honda, M., Saavedra, J. M., & Aguilera, G. (2012). Maternal deprivation in rats is associated with corticotrophin-releasing hormone (CRH) promoter hypomethylation and enhances CRH transcriptional responses to stress in adulthood. J Neuroendocrinol, 24(7), 1055-1064. doi: 10.1111/j.1365-2826.2012.02306.x Curley, J. P., Davidson, S., Bateson, P., & Champagne, F. A. (2009). Social enrichment during postnatal development induces transgenerational effects on emotional and reproductive behavior in mice. Front Behav Neurosci, 3, 25. doi: 10.3389/neuro.08.025.2009 D’Andrea, I., Gracci, F., Alleva, E., & Branchi, I. (2010). Early social enrichment provided by communal nest increases resilience to depression-like behavior more in female than in male mice. Behav Brain Res, 215(1), 71-76. doi: 10.1016/j.bbr.2010.06.030 Francis, D. D., Diorio, J., Plotsky, P. M., & Meaney, M. J. (2002). Environmental enrichment reverses the effects of maternal separation on stress reactivity. J Neurosci, 22(18), 7840-7843. Gottlicher, M. (2004). Valproic acid: an old drug newly discovered as inhibitor of histone deacetylases. Ann Hematol, 83 Suppl 1, S91-92. doi: 10.1007/s00277-004-0850-2 Grayson, D. R., Kundakovic, M., & Sharma, R. P. (2010). Is there a future for histone deacetylase inhibitors in the pharmacotherapy of psychiatric disorders? Mol Pharmacol, 77(2), 126-135. doi: 10.1124/mol.109.061333 Guo, J. U., Su, Y., Zhong, C., Ming, G. L., & Song, H. (2011). Hydroxylation of 5-methylcytosine by TET1 promotes active DNA demethylation in the adult brain. Cell, 145(3), 423-434. doi: 10.1016/j.cell.2011.03.022 Handel, A. E., & Ramagopalan, S. V. (2010). Is Lamarckian evolution relevant to medicine? BMC Med Genet, 11, 73. doi: 10.1186/1471-2350-11-73 Haramati, S., Navon, I., Issler, O., Ezra-Nevo, G., Gil, S., Zwang, R., . . . Chen, A. (2011). MicroRNA as repressors of stress-induced anxiety: the case of amygdalar miR-34. J Neurosci, 31(40), 14191-14203. doi: 10.1523/jneurosci.1673-11.2011 Jenuwein, T., & Allis, C. D. (2001). Translating the histone code. Science, 293(5532), 1074-1080. doi: 10.1126/science.1063127 Jones, P. A. (2012). Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet, 13(7), 484-492. doi: 10.1038/nrg3230 Katada, S., Imhof, A., & Sassone-Corsi, P. (2012). Connecting threads: epigenetics and metabolism. Cell, 148(1-2), 24-28. doi: 10.1016/j.cell.2012.01.001 Kuzumaki, N., Ikegami, D., Tamura, R., Hareyama, N., Imai, S., Narita, M., . . . Suzuki, T. (2011). Hippocampal epigenetic modification at the brain-derived neurotrophic factor gene induced by an enriched environment. Hippocampus, 21(2), 127-132. doi: 10.1002/hipo.20775 Labonte, B., Suderman, M., Maussion, G., Navaro, L., Yerko, V., Mahar, I., . . . Turecki, G. (2012). Genome-wide epigenetic regulation by early-life trauma. Arch Gen Psychiatry, 69(7), 722-731. doi: 10.1001/archgenpsychiatry.2011.2287 Liu, D., Diorio, J., Tannenbaum, B., Caldji, C., Francis, D., Freedman, A., . . . Meaney, M. J. (1997). Maternal care, hippocampal glucocorticoid receptors, and hypothalamic-pituitary-adrenal responses to stress. Science, 277(5332), 1659-1662. Lu, Y., Christian, K., & Lu, B. (2008). BDNF: a key regulator for protein synthesis-dependent LTP and long-term memory? Neurobiol Learn Mem, 89(3), 312-323. doi: 10.1016/j.nlm.2007.08.018 Minichiello, L. (2009). TrkB signalling pathways in LTP and learning. Nat Rev Neurosci, 10(12), 850-860. doi: 10.1038/nrn2738 Morley-Fletcher, S., Rea, M., Maccari, S., & Laviola, G. (2003). Environmental enrichment during adolescence reverses the effects of prenatal stress on play behaviour and HPA axis reactivity in rats. Eur J Neurosci, 18(12), 3367-3374. Mueller, B. R., & Bale, T. L. (2008). Sex-specific programming of offspring emotionality after stress early in pregnancy. J Neurosci, 28(36), 9055-9065. doi: 10.1523/jneurosci.1424-08.2008 Murgatroyd, C., Patchev, A. V., Wu, Y., Micale, V., Bockmuhl, Y., Fischer, D., . . . Spengler, D. (2009). Dynamic DNA methylation programs persistent adverse effects of early-life stress. Nat Neurosci, 12(12), 1559-1566. doi: 10.1038/nn.2436 Musumeci, G., & Minichiello, L. (2011). BDNF-TrkB signalling in fear learning: from genetics to neural networks. Rev Neurosci, 22(3), 303-315. doi: 10.1515/rns.2011.031 Mychasiuk, R., Zahir, S., Schmold, N., Ilnytskyy, S., Kovalchuk, O., & Gibb, R. (2012). Parental enrichment and offspring development: modifications to brain, behavior and the epigenome. Behav Brain Res, 228(2), 294-298. doi: 10.1016/j.bbr.2011.11.036 Nasca, C., Xenos, D., Barone, Y., Caruso, A., Scaccianoce, S., Matrisciano, F., . . . Nicoletti, F. (2013). L-acetylcarnitine causes rapid antidepressant effects through the epigenetic induction of mGlu2 receptors. Proc Natl Acad Sci U S A, 110(12), 4804-4809. doi: 10.1073/pnas.1216100110 Naumova, O. Y., Lee, M., Koposov, R., Szyf, M., Dozier, M., & Grigorenko, E. L. (2012). Differential patterns of whole-genome DNA methylation in institutionalized children and children raised by their biological parents. Dev Psychopathol, 24(1), 143-155. doi: 10.1017/s0954579411000605 Nithianantharajah, J., & Hannan, A. J. (2006). Enriched environments, experience-dependent plasticity and disorders of the nervous system. Nat Rev Neurosci, 7(9), 697-709. doi: 10.1038/nrn1970 Oberlander, T. F., Weinberg, J., Papsdorf, M., Grunau, R., Misri, S., & Devlin, A. M. (2008). Prenatal exposure to maternal depression, neonatal methylation of human glucocorticoid receptor gene (NR3C1) and infant cortisol stress responses. Epigenetics, 3(2), 97-106. Ooi, S. K., Qiu, C., Bernstein, E., Li, K., Jia, D., Yang, Z., . . . Bestor, T. H. (2007). DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature, 448(7154), 714-717. doi: 10.1038/nature05987 Pariante, C. M., & Lightman, S. L. (2008). The HPA axis in major depression: classical theories and new developments. Trends Neurosci, 31(9), 464-468. doi: 10.1016/j.tins.2008.06.006 Roth, T. L., Lubin, F. D., Funk, A. J., & Sweatt, J. D. (2009). Lasting epigenetic influence of early-life adversity on the BDNF gene. Biol Psychiatry, 65(9), 760-769. Roth, T. L., Zoladz, P. R., Sweatt, J. D., & Diamond, D. M. (2011). Epigenetic modification of hippocampal Bdnf DNA in adult rats in an animal model of post-traumatic stress disorder. J Psychiatr Res, 45(7), 919-926. doi: 10.1016/j.jpsychires.2011.01.013 Sapolsky, R. M., Meaney, M. J., & McEwen, B. S. (1985). The development of the glucocorticoid receptor system in the rat limbic brain. III. Negative-feedback regulation. Brain Res, 350(1-2), 169-173. Sato, F., Tsuchiya, S., Meltzer, S. J., & Shimizu, K. (2011). MicroRNAs and epigenetics. FEBS J, 278(10), 1598-1609. doi: 10.1111/j.1742-4658.2011.08089.x Schlinzig, T., Johansson, S., Gunnar, A., Ekstrom, T. J., & Norman, M. (2009). Epigenetic modulation at birth – altered DNA-methylation in white blood cells after Caesarean section. Acta Paediatr, 98(7), 1096-1099. doi: 10.1111/j.1651-2227.2009.01371.x Suri, D., Veenit, V., Sarkar, A., Thiagarajan, D., Kumar, A., Nestler, E. J., . . . Vaidya, V. A. (2012). Early Stress Evokes Age-Dependent Biphasic Changes in Hippocampal Neurogenesis, Bdnf Expression, and Cognition. Biol Psychiatry. doi: 10.1016/j.biopsych.2012.10.023 Szulwach, K. E., Li, X., Li, Y., Song, C. X., Wu, H., Dai, Q., . . . Jin, P. (2011). 5-hmC-mediated epigenetic dynamics during postnatal neurodevelopment and aging. Nat Neurosci, 14(12), 1607-1616. doi: 10.1038/nn.2959 Tsankova, N., Renthal, W., Kumar, A., & Nestler, E. J. (2007). Epigenetic regulation in psychiatric disorders. Nat Rev Neurosci, 8(5), 355-367. doi: 10.1038/nrn2132 Tsankova, N. M., Berton, O., Renthal, W., Kumar, A., Neve, R. L., & Nestler, E. J. (2006). Sustained hippocampal chromatin regulation in a mouse model of depression and antidepressant action. Nat Neurosci, 9(4), 519-525. doi: 10.1038/nn1659 Uddin, M., Aiello, A. E., Wildman, D. E., Koenen, K. C., Pawelec, G., de Los Santos, R., . . . Galea, S. (2010). Epigenetic and immune function profiles associated with posttraumatic stress disorder. Proc Natl Acad Sci U S A, 107(20), 9470-9475. doi: 10.1073/pnas.0910794107 van der Horst, F. C., & van der Veer, R. (2008). Loneliness in infancy: Harry Harlow, John Bowlby and issues of separation. Integr Psychol Behav Sci, 42(4), 325-335. doi: 10.1007/s12124-008-9071-x Weaver, I. C., Cervoni, N., Champagne, F. A., D’Alessio, A. C., Sharma, S., Seckl, J. R., . . . Meaney, M. J. (2004). Epigenetic programming by maternal behavior. Nat Neurosci, 7(8), 847-854. doi: 10.1038/nn1276 Weaver, I. C., Champagne, F. A., Brown, S. E., Dymov, S., Sharma, S., Meaney, M. J., & Szyf, M. (2005). Reversal of maternal programming of stress responses in adult offspring through methyl supplementation: altering epigenetic marking later in life. J Neurosci, 25(47), 11045-11054. doi: 10.1523/jneurosci.3652-05.2005 Weaver, I. C., Diorio, J., Seckl, J. R., Szyf, M., & Meaney, M. J. (2004). Early environmental regulation of hippocampal glucocorticoid receptor gene expression: characterization of intracellular mediators and potential genomic target sites. Ann N Y Acad Sci, 1024, 182-212. doi: 10.1196/annals.1321.099 Weaver, I. C., Meaney, M. J., & Szyf, M. (2006). Maternal care effects on the hippocampal transcriptome and anxiety-mediated behaviors in the offspring that are reversible in adulthood. Proc Natl Acad Sci U S A, 103(9), 3480-3485. doi: 10.1073/pnas.0507526103 Whittle, N., Schmuckermair, C., Gunduz Cinar, O., Hauschild, M., Ferraguti, F., Holmes, A., & Singewald, N. (2013). Deep brain stimulation, histone deacetylase inhibitors and glutamatergic drugs rescue resistance to fear extinction in a genetic mouse model. Neuropharmacology, 64, 414-423. doi: 10.1016/j.neuropharm.2012.06.001 Yehuda, R., Bell, A., Bierer, L. M., & Schmeidler, J. (2008). Maternal, not paternal, PTSD is related to increased risk for PTSD in offspring of Holocaust survivors. J Psychiatr Res, 42(13), 1104-1111. doi: 10.1016/j.jpsychires.2008.01.002 Yehuda, R., & Bierer, L. M. (2009). The relevance of epigenetics to PTSD: implications for the DSM-V. J Trauma Stress, 22(5), 427-434. doi: 10.1002/jts.20448
The Dogma-defying Discovery That Genes Learn From Experience
Haley Peckham.
MA (Philosophy), MSc (Neuroscience) doi: 10.12744/ijnpt.2013.0009-0020
Introduction
Epigenetics challenges our acceptance of the dichotomy between nature and nurture. It is coming to be understood that nurture somehow writes its way into our genes; that experiences, or the effects of experiences on gene transcription, may continue throughout our own lives and into those of the next generation. For anyone engaged in the challenge of helping clients who have suffered traumatic, neglectful or abusive experiences, or for anyone offering clients a new experience through the psychotherapeutic relationship, an understanding of epigenetics is pertinent and illuminating. Here I introduce the concept of epigenetics, describe some epigenetic mechanisms, and review the findings of a number of recent studies that seek to understand why, what, and how our genes can learn from experience. The engaging debate of whether it is nature or nurture that has the most influence over who we are and what kind of lives we lead often illuminates the attitudes the debaters hold with regard to their own lives. In my early twenties, I tended to champion the influence of the environment, as although I felt affected by childhood adversity, I did not like to think of myself as passively, fatalistically accepting nature—my genetic inheritance—as my lot in life. Discoveries in biology have made this debate and my wrangling with it entirely passé with the enthralling field of epigenetics, which allows for degrees of gene expression or “shades of grey” that release us from the “black and white” dichotomy of genetic determinism. Epigenetics provides insights into how the environment dynamically impacts on our gene expression as humans, influencing, amongst other traits, our and our children’s health, ability to learn and remember, and responses to stress. The fabric of the lives led by our parents and grandparents, from their diet (Katada, Imhof, & Sassone-Corsi, 2012) to their education, the care they received, traumas they may have suffered, and perhaps many other experiences leave their legacy written alongside our DNA as “instructions for interpretation.” The purpose of this as yet inappreciably sophisticated and elegant code of epigenetic information, if it can be properly be called a “purpose,” is to record, use, and pass on information about the intricacies of the environment experienced in both our lives and those that have gone before us, so that we and our children can be as well prepared for survival in our particular environment as it possible to be. Life, over lives, continually perceives the environment, and the epigenetic instructions for interpreting the DNA are written, used, revised, and may be inherited, along with our genes. Epigenetics is the contemporary study of how the environment influences gene expression both within and, through heritable changes in DNA, beyond the lifetime of an organism. The idea that organisms can inherit environmentally acquired characteristics is, however, the old idea of Lamarckian inheritance. In 1809, Jean-Baptiste Lamarck suggested that an organism would acquire traits through adapting to its environment, and that these traits would then be inherited by its offspring (Handel & Ramagopalan, 2010). Lamarck’s theory was overlooked in favor of Darwin’s natural selection theory of evolution, as the two explanations appeared at the time to be mutually exclusive, but the advent of epigenetics has made it possible for these theories to be reconciled.
“Epigenetics” literally means “above the genes,” and is the means by which the environment “marks” the genes, dramatically or subtly, changing their level of expression either transiently or for our lifetime, or, through inheritability, throughout our children’s and grandchildren’s lifetimes. A formal definition of epigenetic events proposed by Adrian Bird is: “the structural adaption of chromosomal regions so as to register, signal or perpetuate altered activity states” (Bird, 2007, p. 398). This definition encompasses the broad remit of epigenetic marks from transient, where the epigenetic mark ascribed by the environmental adaption lasts only a few hours, to heritable, where the environmental effects last over a generation. The brain-derived neurotrophic factor (BDNF) gene, implicated in psychiatric disorders and learning and memory (Autry & Monteggia, 2012; Boulle et al., 2012; Lu, Christian, & Lu, 2008; Minichiello, 2009; Musumeci & Minichiello, 2011), is subject to both short- and long-term epigenetic marking in rodents. For example, following the favorable social experience of being reared in a communal nest, mice challenged with one hour in a mildly stressful novel environment generate hippocampal BDNF faster than mice raised in a standard nest, as a result of an epigenetic mark on the BDNF gene (Branchi, Karpova, D’Andrea, Castren, & Alleva, 2011). However, rat pups subjected to a rat equivalent of childhood maltreatment are epigenetically marked by this experience, reducing the level of BDNF in their pre-frontal cortex throughout their adult life. The offspring of these rats also carry the same epigenetic mark on their BDNF gene even when they have been cross-fostered to non-maltreating mothers (Roth, Lubin, Funk, & Sweatt, 2009). Thus, epigenetics, the mechanisms by which our genes record or adapt to the environment, can shape gene expression over a few minutes, an hour, or a lifetime, and can even shape the gene expression pattern of the next generation. It is even possible for genes to “remember” an event and make a contingency plan for its recurrence, as in the case of the corticotropin-releasing hormone gene of rat pups. In response to maternal deprivation, the promoter region of this gene is epigenetically marked. Later, following a stressful experience in adulthood, the pre-recorded epigenetic mark leads to a hypersensitive stress response, observed as a more actively transcribed corticotropin-releasing hormone gene and increased levels of the stress hormone corticosterone (Chen et al., 2012). There are three main types of epigenetic modification: DNA methylation and histone modification, outlined here together with examples of these mechanisms in action, and translational regulation by micro RNA (not pertinent to this review, but see (Haramati et al., 2011; Sato, Tsuchiya, Meltzer, & Shimizu, 2011) for useful introductions). Types of epigenetic modification, Level I: DNA methylation Methylation involves the addition of a methyl group to mammalian DNA, and can occur in response to environmental influence, making a stable, potentially heritable addition to the DNA that enhances or represses the transcription of a gene. Methylation is not a mutation, as the sequence of bases (adenine, guanine, cytosine, and thymine) remains the same. The methyl group is added to a cytosine nucleotide, usually followed by a guanine nucleotide (a CpG site), by enzymes known as DNA methyltransferases (DNMTs). The promoter regions or transcriptional start site of genes (the point at which RNA polymerase II begins transcription of the DNA into mRNA) are frequently rich in CpG sites, and methylation of these regions is associated with long-term silencing of genes. Methylation of CpG sites within the body of the gene is more ambiguous and context specific and can lead to either repression or activation (Jones, 2012)’. Methylated DNA can also be bound by Methyl Binding Domain (MBD) proteins such as MeCP2 which recruit other protein complexes to re-model the local chromatin (DNA + histone packaging), leading either to repression or activation of specific genes. MeCP2 is an important point of integration between types of epigenetic modification, as MeCP2 binds methylated DNA but also can recruit chromatin re-modeling proteins which implement the other major type of epigenetic alteration, histone modification, described below (Cedar & Bergman, 2012). Methylation of DNA is reversible but instead of de-methylation occurring via the action of a single enzyme, it occurs through a multistep process. It is intriguing that almost half of the methylated DNA in the brain is 5-hydroxymethylcytosine, an intermediate formed during the multi-step de-methylation of DNA (Szulwach et al., 2011), a process which is dependent on neuronal activity (Guo, Su, Zhong, Ming, & Song, 2011). The contemporary view of DNA methylation is that, in the brain especially, it is a highly dynamic process, responsive to neural activity (and therefore experience) and crucial for plasticity-related functions such as learning, memory and behavioral adaptation (Baker-Andresen, Ratnu, & Bredy, 2013). Types of epigenetic modification, Level II: Histone modification The DNA in the nucleus of each eukaryotic cell, including in the case of humans, contains all of the genetic material. So a human skin cell carries the same complete genetic code as a neuron or a parietal cell of the stomach. Each specialized cell need only be able to transcribe a small fraction of the total DNA it carries in order to perform its specific function. Since the rest of its DNA will never need to be accessed, it can be packed away as efficiently as possible. Histone proteins facilitate this efficient packaging of DNA, largely due to the electrostatic attraction between the negatively charged DNA and the positively charged histone proteins. DNA is coiled around a core octamer of histone proteins, like thread on a spool, forming a single nucleosome. Multiple nucleosomes together are known as chromatin, which has different conformations reflecting the accessibility of the genes for transcription. Euchromatin is open chromatin, conceivable as separate beads on a string; here the genes are accessible, whereas in condensed chromatin or heterochromatin, the genes are inaccessible and un-transcribed, as they are so densely packed (Jenuwein & Allis, 2001).
Figure 1. Structure of a nucleosome: DNA wound around a histone core. From “Epigenetic Regulation in Psychiatric Disorders,” by N. Tsankova, W. Renthal, A. Kumar, and E. J. Nestler, 2007, Nature Reviews Neuroscience, 8, p. 356. Copyright 2007 by Nature Publishing Group. Reproduced with permission. Like most proteins, the histone proteins comprising the core around which the DNA winds itself can have chemical groups attached to them that may either subtly change their charge, and thus how tightly they are bound to the DNA, or change their affinity for other protein-binding partners. In this way, histone modifications alter the chromatin structure and hence the accessibility of genes, but can also affect the binding of other molecules including DNA methyltransferases—the enzymes that add methyl groups to DNA, not the histone proteins that the DNA coils around (Ooi et al., 2007). This interaction forms another point of integration between DNA-modifying and histone-modifying epigenetic mechanisms. Histones may be acetylated, methylated (including bi- or tri-methylated), phosphorylated or ubiquitinylated on specific residues, and each group or combination of groups may signal a precise outcome. For example, an epigenetic modification of a tri-methylated lysine4 residue on histone3 signals an actively transcribed gene, whereas a tri-methylation on lysine27 of histone3 signals a repressed gene state. If lysines 4 and 27 are both tri-methylated, the gene is thought to be “poised,” ready for activation (Bernstein et al., 2006). As there are four histones (two of each make up the octamer core), and at least four chemical groups that can attach to multiple residues, there are a large number of possible combinations of epigenetic marks that can be made on a histone octamer, giving rise to the concept of a “histone code” that can nuance gene expression (Jenuwein & Allis, 2001). There are enzymes that catalyze the addition and removal of each type of chemical group. Histone acetylases (HATs), for example, add acetyl groups to the histone, reducing the electrostatic charge difference and therefore the affinity between the histone and the DNA, thus loosening the coil. Accordingly, HATs tend to be associated with gene activation, whereas histone de-acetylases (HDACs) remove the acetyl group, increase the electrostatic attraction, tighten the coil, and are thus considered transcriptional repressors (Bannister & Kouzarides, 2011). In contrast to the epigenetic modification of a methylation made to DNA, histone modifications are easily reversible and highly dynamic, although the enzymes that catalyze the addition and removal of various groups are themselves regulated. The histone code also has great flexibility in determining how long a gene’s activity may be enhanced or repressed, as epigenetic marks may record an experience on a gene without necessarily changing its level of transcription. They may transiently enhance or repress gene expression, or may recruit (or repel) DNA methylases that can administer de novo methylations on the DNA which result in the gene becoming stably repressed throughout the life of the organism and in the next generation (Ooi, et al., 2007). It is interesting to note that Valproic acid, a known teratogen used in the treatment of bipolar disorder, is a histone de-acetylase inhibitor but also leads to reduced levels of DNA methylation, demonstrating that a transient and easily reversible epigenetic histone acetylation is frequently followed by a more stable and less easily reversed DNA methylation, silencing the gene (Alonso-Aperte, Ubeda, Achon, Perez-Miguelsanz, & Varela-Moreiras, 1999; Gottlicher, 2004). Thus, histone modification and DNA methylation, whilst distinct epigenetic mechanisms, can act in synergy to produce permanent gene silencing. Figure 2. The dynamic addition and removal of chemical groups to histone tails by specific enzymes. From “Epigenetic Regulation in Psychiatric Disorders,” by N. Tsankova, W. Renthal, A. Kumar, and E. J. Nestler, 2007, Nature Reviews Neuroscience, 8, p. 356. Copyright 2007 by Nature Publishing Group. Reproduced with permission. The epigenetic impact of early maternal care Many animal studies of epigenetics have looked at the impact of early maternal care on the hypothalamic-pituitary-adrenal (HPA) axis or, put more simply, the stress axis. This axis co-ordinates the response to stress from the brain, which perceives the stress, to the adrenal gland, which releases glucocorticoids such as cortisol (corticosterone in rodents) in response to the stressor. The hypothalamus receives neural input—our perception of the stress—and releases corticotropin-releasing hormone (CRH) that acts on the anterior pituitary gland to release adrenocorticotropic hormone (ACTH). This in turn acts on the cortex of the adrenal gland to release cortisol. The multiple steps in this process allow multiple opportunities for the glucocorticoids to negatively feed back at several levels of the HPA axis, including the hippocampus and the hypothalamus, limiting CRH release, and at the pituitary, limiting ACTH release (Anacker, Zunszain, Carvalho, & Pariante, 2011; Pariante & Lightman, 2008). Glucocorticoids feed back via glucocorticoid receptors. More receptors means a swifter, more efficient negative feedback mechanism resulting in lower levels of circulating CRH, ACTH and cortisol, and consequently an HPA axis that readily returns to homeostasis. However, in response to chronically high levels of circulating glucocorticoids, glucocorticoid receptors are down-regulated; as a result, the negative feedback mechanism is less efficient, and the HPA axis is slower to return to homeostasis or may become dysregulated (Anacker, et al., 2011; Sapolsky, Meaney, & McEwen, 1985). The HPA axis is an adaptive system subject to early-life epigenetic programming both prenatally and postnatally through variations in maternal care, separation and abuse. Early pre-natal stress in mice has been epigenetically linked to enduring changes in HPA axis reactivity and a depressive phenotype in male offspring. Male adult mice whose mothers had been mildly stressed early in utero had higher levels of CRH and fewer glucocorticoid receptors in their hippocampus, the former increasing the activation of the HPA axis and the latter reducing the ability to regulate the HPA axis. These mice exhibited behaviors associated with depression and, following exposure to restraint stress, had increased levels of corticosterone compared to mice whose mothers had not been subjected to early pre-natal stress. When the promoter regions of relevant genes were examined, it was discovered that the promoter for corticotropin-releasing hormone was hypo-methylated, increasing the expression of CRH, the hormonal activator of the HPA axis. The glucocorticoid receptor promoter was also hyper-methylated, reducing its level of expression and thereby the mouse’s ability to negatively regulate its HPA axis after activation by CRF. This would at least contribute to, if not cause, the observed depressed behavior (Mueller & Bale, 2008). Figure 3. Combinations of histone marks corresponding to active, permissive, repressed or inactive (silenced) genes. From “Epigenetic Regulation in Psychiatric Disorders,” by N. Tsankova, W. Renthal, A. Kumar, and E. J. Nestler, 2007, Nature Reviews Neuroscience, 8, p. 356. Copyright 2007 by Nature Publishing Group. Reproduced with permission. Early post-natal stress has also been linked to enduring epigenetic changes that alter the reactivity of the HPA axis. Rat pups that receive enhanced licking and grooming by their mothers in the first ten days of life acquire a permanent increase in their number of glucocorticoid receptors, meaning that their HPA axis is more readily negatively regulated. Thus, when these pups become adults and are exposed to an acute stressor, they release corticosterone, which negatively feeds back through a greater number of glucocorticoid receptors, quickly reducing the levels of CRH, circulating ACTH, and corticosterone (Liu et al., 1997). If this effect was correlated with measurable changes to the epigenome, i.e., DNA methylation or histone code changes at the site of a relevant gene, it would support the hypothesis that maternal care leads to epigenetic changes that moderate the activity of the stress axis. In support of this hypothesis, a landmark study by Weaver’s group (Weaver, Diorio, Seckl, Szyf, & Meaney, 2004) compared rat pups that had either high or low levels of licking and grooming from mothers with whom they were cross-fostered, and found epigenetic differences in the promoter region of the glucocorticoid receptor gene. Pups that had received high levels of licking and grooming had hypo-methylated DNA and hyper-acetylated histone3 at the promoter region of the glucocorticoid receptor gene where a transcription factor binds, leading to enhanced transcriptional activity of the glucocorticoid receptor gene and therefore to more glucocorticoid receptors. As adults, pups that had received low levels of licking and grooming had fewer glucocorticoid receptors and correspondingly higher levels of circulating corticosterone following restraint stress. Taking the study a step further towards identifying maternal care as a cause of epigenetic changes that modify the reactivity of the HPA axis, a histone de-acetylase (HDAC) inhibitor was administered to the rats which not only enhanced acetylation of the glucocorticoid promoter in the low licking and grooming group, it also removed all of the other observed differences in this group: the glucocorticoid promoter became hypo-methylated, more glucocorticoid receptors were expressed, and the corticosterone response following restraint stress was normalized to the same level as the high licking and grooming group. Essentially, the epigenetic differences brought about by lower levels of maternal care were reversed by the HDAC inhibitor (Weaver et al., 2004). In a follow-up study by the same group (Weaver et al., 2005), rats were infused with a methyl group donator. Even if the rats had received high levels of licking and grooming by their mothers when they were pups, leading to hypo-methylation of the glucocorticoid promoters, if they received the methylating drug as adults, all of the beneficial effects of this extra licking and grooming were reversed; the glucocorticoid promoter became hyper-methylated and fewer glucocorticoid receptors were expressed, leading to higher levels of corticosterone following a stressful experience and higher scores in a test of behavioral responses to stress (Weaver et al., 2005). From their combined studies, the authors primarily concluded that maternal behavior programs the stress response through epigenetic modification of the glucocorticoid promoter, and secondarily concluded that although maternal care makes stable changes to the epigenome, these are pharmacologically modifiable even in adults (Weaver, et al., 2005). Since then, in the case of studies showing that differences in estrogen receptor levels correlated with the level of maternal care received as a pup and the level of maternal care given in turn to offspring (Champagne, Weaver, Diorio, Sharma, & Meaney, 2003), the relationship has also been found to be mediated through epigenetic changes to the estrogen receptor promoter. Thus, receiving a high level of maternal care leads to reduced methylation of the CpG region within the promoter of the estrogen receptor, correspondingly increased levels of expressed estrogen receptors, and the capacity to give a high level of maternal care to the following generation of pups (Champagne et al., 2006). Separation from mother in infancy has long been known to have emotionally and psychologically detrimental sequelae, and the work of two brilliant and influential researchers, Harry Harlow and John Bowlby, is eloquently discussed in van der Horst and van der Veer’s review (van der Horst & van der Veer, 2008). In the field of epigenetics, the effects of maternal separation on the reactivity of the HPA axis are beginning to be discovered. In the HPA axis, stressful stimuli perceived at the level of the hypothalamus lead to secretion of CRH-stimulating ACTH. Arginine vasopressin (AVP) is also secreted by the hypothalamus, and potentiates the effect of CRH at the anterior pituitary, stimulating more ACTH release and increased HPA activity. Mice that were separated from their mothers for three hours a day for the first ten days of their lives had higher basal levels of corticosterone as well as higher levels following a stressful experience, and also scored higher on behavioral measures of stress and lower on memory tasks. Higher levels of expressed AVP correlated with hypo-methylated DNA in the regulatory region of the AVP gene (Murgatroyd et al., 2009). The methyl-binding protein MeCP2, which binds methylated DNA, ordinarily represses AVP transcription. But in mice that have been separated from their mothers, MeCP2 cannot bind as readily to the AVP promoter, as it is hypo-methylated even when the mouse reaches adulthood, leading to reduced repression by MeCP2 and consequent higher levels of AVP transcription. This results in a persistent hormonal activation of the HPA axis throughout the life of the mouse (Murgatroyd, et al., 2009). Figure 4. The hypothalamic pituitary adrenal axis and its regulation by glucocorticoid negative feedback. Illustration by Brian M. Sweis (2012). Reproduced from http://commons.wikimedia.org/wiki/File:HPA_Axis_Diagram_(Brian_M_Sweis_2012).png As well as enduringly activating the HPA axis, early life stress induced by maternal separation has been shown to enhance performance in stress-related memory tasks in young rats (Suri et al., 2012). The rats’ improved performance correlated with increased neurogenesis, reduced repressive histone methylation of a BDNF gene promoter, and correspondingly higher levels of BDNF in the hippocampus, compared to controls. Interestingly, once the maternally separated rats reached middle age, all these beneficial effects were lost, although this could be ameliorated by long-term antidepressant treatment. The authors of this study suggested that maternal separation stress induces biologically adaptive responses including epigenetic changes that increase the rat pups’ chances of having to survive alone, without the mother’s help and protection, but that these adaptive changes exact a heavy toll when the rat reaches middle age (Suri, et al., 2012). From an evolutionary point of view, as long as the adaptive response to maternal separation allows the rat to survive on its own long enough to be able to reproduce, the exacted toll is worthwhile. Paradigms that mimic abusive behavior in humans have also been used to examine the epigenomic effects of maternal abuse in rodents, demonstrating that abuse leads to significant hypo-methylation of the BDNF promoter in the pre-frontal cortex. Notably, rats that had suffered maltreatment in infancy also maltreated their own (or cross-fostered) pups, and the effect of their own maltreatment could be seen in their biological offspring as a hypo-methylated BDNF promoter region even when the pups were fostered to a caring mother (Roth, et al., 2009). Maltreatment, or abuse, appears to be such a profound environmental influence that its effects linger in the generation following that which experienced the maltreatment firsthand. A similar observation of the trans-generational transmittance of a profound experience has been made by Rachel Yehuda, a researcher into post-traumatic stress disorder (PTSD) who found a higher than normal prevalence of PTSD in the children of mothers who had PTSD as a result of the Holocaust, suggesting a mechanism other than traditional genetic contribution in play, with epigenetics being the obvious candidate (Yehuda, Bell, Bierer, & Schmeidler, 2008). Although specific epigenetic modifications have not yet been linked to experiences that lead to the development of PTSD in humans, a recent study has identified patterns of genes that are differentially methylated in human sufferers of PTSD (Uddin et al., 2010). Additionally, in the rat model of PTSD, where the animals are exposed on two occasions to a cat, as well as having a new rat cage mate daily for 31 days, dramatic epigenetic changes have been observed in the promoter region of the BDNF gene in the hippocampus (Roth, Zoladz, Sweatt, & Diamond, 2011). Taken together, these results suggest that the development of human PTSD may occur through traumatic events causing epigenetic changes to specific genes, which, as well as leading to PTSD in the trauma victim, could confer an increased risk of the same in the victim’s children, as the epigenetic signatures on the genes caused by the trauma could be inherited (Yehuda & Bierer, 2009). Although speculative, it seems likely that the inheritance of genes that have been epigenetically marked by trauma could be biologically adaptive and enhance the chance of survival if the environment remains life-threatening for the next generation. Hyper-vigilance and extreme alertness following trauma, whilst profoundly exhausting and distressing, may be life-saving in a dangerous environment. Human studies looking at the epigenetic impact of the environment Gathering data for human studies tends to involve the use of postmortem tissue and detailed retrospective analyses of significant life events or trauma. A recent study using these methodologies compared the differences in DNA methylation across the whole genome of 25 men with a history of severe child abuse compared to 16 controls, and found that fewer genes are actively transcribed in men who suffered abusive childhoods. The methylation profiles of the men showed 248 genes were hyper-methylated and 114 genes were hypo-methylated in the group with a history of abuse compared to the control group (Labonte et al., 2012). Of these, the top five most hyper-methylated genes were neuronal, and many of the genes that were differently methylated in the group who had experienced childhood abuse were related to plasticity, i.e., genes that are known to have a role in learning or adaptive mechanisms. A similar study using blood samples found that children raised in institutions had more methylated, and thus fewer expressed, genes than children raised by their biological parents (Naumova et al., 2012). It is tempting to speculate that perhaps lower levels of care restrict the variety of genes that can be expressed, whereas higher levels of care are the epigenetic gateway to our available genome. We speak of wanting to give our children “every opportunity”, but the reality of what epigenetic opportunities are available to our children could largely depend on the care we give them very early in life. Access to the widest potential of their genome could be profoundly influenced by their experience of our care, and if the genes that are accessed by enhanced care perform plasticity-related functions, the effect of that care is compounded for good or ill. Poor care may mean that fewer plasticity-related genes are expressed, thus restricting a child (and the adult’s) potential to learn, remember or adapt within their environment, manifesting in less flexibility and perhaps more stereotyped or rigid responses. Studies that identify the epigenetic effects of levels of care or experiences on specific genes have been undertaken in humans. In a striking human parallel to Weaver’s (2004) rodent study showing that low levels of maternal care epigenetically modified the glucocorticoid receptor promoter by hyper-methylation, reducing the transcription of the glucocorticoid receptor gene, McGowan et al. (2009) found that suicide victims who had a history of childhood abuse had hyper-methylated glucocorticoid promoters and fewer HPA axis-regulating hippocampal glucocorticoid receptors than controls. Both prenatal stress and even the birth experience have also been linked to epigenetic changes. The depressive mood of mothers in the third trimester of pregnancy has been linked to hyper-methylated glucocorticoid receptor promoters and higher cortisol responses in three-month-old babies (Oberlander et al., 2008), while infants born by Caesarean section have significantly higher levels of DNA methylation in their leucocytes compared to vaginal births at the time of delivery, although this normalizes after 3–5 days (Schlinzig, Johansson, Gunnar, Ekstrom, & Norman, 2009).
Pharmacotherapeutics, enriching environments, and reversible epigenetic change
Pharmacological treatments can reverse epigenetic marks caused by environmental experiences on both the BDNF gene (Roth, et al., 2009; Suri, et al., 2012) and the glucocorticoid receptor gene (Weaver, et al., 2005; Weaver, Meaney, & Szyf, 2006). Antidepressant treatment has frequently been shown to restore levels of BDNF in animal models of depression (Balaratnasingam & Janca, 2012), and in the social defeat, mouse model of depression, imipramine has been demonstrated to act through epigenetic mechanisms, reversing the down-regulating effect that social defeat has on BDNF transcription levels, although it does so by a compensatory HDAC-inhibiting mechanism rather than direct reversal of repressive histone methylations (N. M. Tsankova et al., 2006). The HDAC inhibitor, valproic acid, a treatment in certain psychiatric disorders, also epigenetically enhances BDNF transcription, facilitating the forgetting (or extinction) of fear-conditioned learning (Bredy et al., 2007; Whittle et al., 2013). Other histone-modifying agents may become useful psychoactive medications. It has recently been demonstrated that the acetylating agent L-acetyl-carnitine, already available as a dietary supplement, enhances transcription of the type II metabotropic glutamate receptor, which has a swift and enduring anti-depressant effect in rodents (Nasca et al., 2013). As epigenetic mechanisms mediate the effects of environment, the involvement of epigenetic mechanisms in psychiatric disorders that have an environmental component is axiomatic. Whilst pharmacotherapeutics holds promise for the treatment of psychiatric disorders (Grayson, Kundakovic, & Sharma, 2010; N. Tsankova, Renthal, Kumar, & Nestler, 2007), for the many people who fall below the threshold of a psychiatric diagnosis or who prefer non-medicating treatments, the very nature of the epigenetic mechanism suggests an alternative to drug treatment in the form of new, desirable environmental experiences to overwrite previous negative or undesirable experiences written on the epigenome. Environmental enrichment is proof of the principle that new environmental experiences can reverse or ameliorate the epigenetic effects of a previously impoverished or stressful environment. Enriching environments provide opportunities for enhanced sensory, motor, cognitive, and social stimulation, and have a multitude of beneficial effects, from enhancing neural plasticity and learning and memory, to conferring resilience to depression and ameliorating the effects of many brain disorders including Alzheimer’s and Huntington’s diseases and stroke (Bekinschtein, Oomen, Saksida, & Bussey, 2011; D’Andrea, Gracci, Alleva, & Branchi, 2010; Nithianantharajah & Hannan, 2006). The effects of an enriched environment in rodents are profound, and can even mitigate the effects of prenatal stress or maternal separation on the HPA axis (Francis, Diorio, Plotsky, & Meaney, 2002; Morley-Fletcher, Rea, Maccari, & Laviola, 2003). Analogously to the effects of trauma, the effects of enrichment can be seen in the next generation, improving learning and memory (Arai, Li, Hartley, & Feig, 2009) and, following the socially enriching experience of being reared in a communal nest, enhancing levels of maternal care in the next generation, reducing anxiety behavior, and increasing litter sizes (Curley, Davidson, Bateson, & Champagne, 2009). It is likely that epigenetic mechanisms mediate the benefits of enrichment in terms of both the generation experiencing the enrichment and the trans-generational effects of enrichment that are observed. Recent work shows that rats whose parents had enriching experiences have less methylated DNA than rats whose parents experienced standard housing (Mychasiuk et al., 2012), suggesting that enrichment (such as high levels of care) increases the number of genes that will be actively transcribed. This perhaps reflects a need for greater biological complexity in order to be able to thrive in a more complex (enriched) environment. One gene that is extremely responsive to the environment is BDNF, which is epigenetically modified by the experience of abuse (see above) as well as by the experience of an enriched environment. A recent study demonstrated increased levels of permissive histone methylations, enhancing transcription of the gene and raising levels of hippocampal BDNF in mice that experienced a month in an enriched environment (Kuzumaki et al., 2011). The role of BDNF in learning and memory behaviors, combined with its epigenetically regulated transcription, suggests this gene is a critical transducer of the experienced environment. Environmental enrichment is a powerful tool to effect epigenomic changes, which subsequently affect the levels of BDNF and modulate HPA axis activity, which, in turn, builds resilience to, or delays the onset of, various psychiatric or neurodegenerative disorders.
The relevance of epigenetics for
psychotherapy
When I learned that real-life subjective experiences could regulate gene transcription, it changed the way I understood myself. I was not just an immutable collection of transcribed genes. My genes had my experiences of early care, my traumas and dramas and education written onto them in the tiniest molecular inscription of the histone code and DNA methylations. Furthermore, even as I was born, the most salient facts of my parents’ and even their parents’ lives—the kinds of care they had received, their education, their sense of safety or fear—was etched into my genes too, to help me adapt to what was biologically anticipated to be a similar environment. I did not know about epigenetics when I underwent psychotherapy, but if I had, it may have inspired my curiosity about my parents’ lives and informed my developing narrative that who they were, who I was, was a more intricate and elegantly entwined interplay between our genes and our environments than I could ever have imagined. Neither they nor I was responsible or to blame; instead, we each were simply trying to adapt to our niche, trying to make the best of what we had: our genes and our unknown, influential, epigenetic legacy. In writing this article I wanted to share the concept of epigenetics and to review and communicate the most relevant findings in contemporary epigenetics research with psychotherapists or anyone who seeks to understand and alleviate emotional and psychological distress.
Epigenetics is an astounding and revolutionary discovery that brings compassionate understanding and great hope for the future to anyone who suffers distress, as well as those who work so intimately with distressed clients. An appreciation of epigenetics inspires compassion for those of us who come into the world vigilant and defensive, our epigenetic legacy anticipating a harsh world, a dangerous place to live; yet it also inspires hope, for even the most epigenetically determined aggressive, defensive stance is, by its very nature, responsive to the novelty of benign and benevolent environments.
Fundamentally, epigenetics translates experience in the world into a gene expression profile that shapes who we are. If we are to develop or change ourselves, we must actively seek the experiences that will help us develop along the trajectory we desire. Psychotherapy can be seen as a form of emotional, environmental enrichment. It is perfectly placed to offer an enriched experience of a caring relationship, albeit an asymmetrical one, where one can received empathic, attuned attention that may be wholly novel and life changing. By offering a new, benevolent experience of relationship, psychotherapy can be reasonably assumed to change gene expression, calming a hyper-reactive stress axis and ameliorating the effects of an epigenetically scarred BDNF promoter, and in so doing, changing our responses to stress and perhaps our capacity to learn and flexibly respond to emotionally challenging situations. For the client in psychotherapy, as in many situations in life, it really is the experience that counts.
References Alonso-Aperte, E., Ubeda, N., Achon, M., Perez-Miguelsanz, J., & Varela-Moreiras, G. (1999). Impaired methionine synthesis and hypomethylation in rats exposed to valproate during gestation. Neurology, 52(4), 750-756. Anacker, C., Zunszain, P. A., Carvalho, L. A., & Pariante, C. M. (2011). The glucocorticoid receptor: pivot of depression and of antidepressant treatment? Psychoneuroendocrinology, 36(3), 415-425. doi: 10.1016/j.psyneuen.2010.03.007 Arai, J. A., Li, S., Hartley, D. M., & Feig, L. A. (2009). Transgenerational rescue of a genetic defect in long-term potentiation and memory formation by juvenile enrichment. J Neurosci, 29(5), 1496-1502. doi: 10.1523/jneurosci.5057-08.2009 Autry, A. E., & Monteggia, L. M. (2012). Brain-derived neurotrophic factor and neuropsychiatric disorders. Pharmacol Rev, 64(2), 238-258. doi: 10.1124/pr.111.005108 Baker-Andresen, D., Ratnu, V. S., & Bredy, T. W. (2013). Dynamic DNA methylation: a prime candidate for genomic metaplasticity and behavioral adaptation. Trends Neurosci, 36(1), 3-13. doi: 10.1016/j.tins.2012.09.003 Balaratnasingam, S., & Janca, A. (2012). Brain Derived Neurotrophic Factor: a novel neurotrophin involved in psychiatric and neurological disorders. Pharmacol Ther, 134(1), 116-124. doi: 10.1016/j.pharmthera.2012.01.006 Bannister, A. J., & Kouzarides, T. (2011). Regulation of chromatin by histone modifications. Cell Res, 21(3), 381-395. doi: 10.1038/cr.2011.22 Bekinschtein, P., Oomen, C. A., Saksida, L. M., & Bussey, T. J. (2011). Effects of environmental enrichment and voluntary exercise on neurogenesis, learning and memory, and pattern separation: BDNF as a critical variable? Semin Cell Dev Biol, 22(5), 536-542. doi: 10.1016/j.semcdb.2011.07.002 Bernstein, B. E., Mikkelsen, T. S., Xie, X., Kamal, M., Huebert, D. J., Cuff, J., . . . Lander, E. S. (2006). A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell, 125(2), 315-326. doi: 10.1016/j.cell.2006.02.041 Bird, A. (2007). Perceptions of epigenetics. Nature, 447(7143), 396-398. doi: 10.1038/nature05913 Boulle, F., van den Hove, D. L., Jakob, S. B., Rutten, B. P., Hamon, M., van Os, J., . . . Kenis, G. (2012). Epigenetic regulation of the BDNF gene: implications for psychiatric disorders. Mol Psychiatry, 17(6), 584-596. doi: 10.1038/mp.2011.107 Branchi, I., Karpova, N. N., D’Andrea, I., Castren, E., & Alleva, E. (2011). Epigenetic modifications induced by early enrichment are associated with changes in timing of induction of BDNF expression. Neurosci Lett, 495(3), 168-172. doi: 10.1016/j.neulet.2011.03.038 Bredy, T. W., Wu, H., Crego, C., Zellhoefer, J., Sun, Y. E., & Barad, M. (2007). Histone modifications around individual BDNF gene promoters in prefrontal cortex are associated with extinction of conditioned fear. Learn Mem, 14(4), 268-276. doi: 10.1101/lm.500907 Cedar, H., & Bergman, Y. (2012). Programming of DNA methylation patterns. Annu Rev Biochem, 81, 97-117. doi: 10.1146/annurev-biochem-052610-091920 Champagne, F. A., Weaver, I. C., Diorio, J., Dymov, S., Szyf, M., & Meaney, M. J. (2006). Maternal care associated with methylation of the estrogen receptor-alpha1b promoter and estrogen receptor-alpha expression in the medial preoptic area of female offspring. Endocrinology, 147(6), 2909-2915. doi: 10.1210/en.2005-1119 Champagne, F. A., Weaver, I. C., Diorio, J., Sharma, S., & Meaney, M. J. (2003). Natural variations in maternal care are associated with estrogen receptor alpha expression and estrogen sensitivity in the medial preoptic area. Endocrinology, 144(11), 4720-4724. doi: 10.1210/en.2003-0564 Chen, J., Evans, A. N., Liu, Y., Honda, M., Saavedra, J. M., & Aguilera, G. (2012). Maternal deprivation in rats is associated with corticotrophin-releasing hormone (CRH) promoter hypomethylation and enhances CRH transcriptional responses to stress in adulthood. J Neuroendocrinol, 24(7), 1055-1064. doi: 10.1111/j.1365-2826.2012.02306.x Curley, J. P., Davidson, S., Bateson, P., & Champagne, F. A. (2009). Social enrichment during postnatal development induces transgenerational effects on emotional and reproductive behavior in mice. Front Behav Neurosci, 3, 25. doi: 10.3389/neuro.08.025.2009 D’Andrea, I., Gracci, F., Alleva, E., & Branchi, I. (2010). Early social enrichment provided by communal nest increases resilience to depression-like behavior more in female than in male mice. Behav Brain Res, 215(1), 71-76. doi: 10.1016/j.bbr.2010.06.030 Francis, D. D., Diorio, J., Plotsky, P. M., & Meaney, M. J. (2002). Environmental enrichment reverses the effects of maternal separation on stress reactivity. J Neurosci, 22(18), 7840-7843. Gottlicher, M. (2004). Valproic acid: an old drug newly discovered as inhibitor of histone deacetylases. Ann Hematol, 83 Suppl 1, S91-92. doi: 10.1007/s00277-004-0850-2 Grayson, D. R., Kundakovic, M., & Sharma, R. P. (2010). Is there a future for histone deacetylase inhibitors in the pharmacotherapy of psychiatric disorders? Mol Pharmacol, 77(2), 126-135. doi: 10.1124/mol.109.061333 Guo, J. U., Su, Y., Zhong, C., Ming, G. L., & Song, H. (2011). Hydroxylation of 5-methylcytosine by TET1 promotes active DNA demethylation in the adult brain. Cell, 145(3), 423-434. doi: 10.1016/j.cell.2011.03.022 Handel, A. E., & Ramagopalan, S. V. (2010). Is Lamarckian evolution relevant to medicine? BMC Med Genet, 11, 73. doi: 10.1186/1471-2350-11-73 Haramati, S., Navon, I., Issler, O., Ezra-Nevo, G., Gil, S., Zwang, R., . . . Chen, A. (2011). MicroRNA as repressors of stress-induced anxiety: the case of amygdalar miR-34. J Neurosci, 31(40), 14191-14203. doi: 10.1523/jneurosci.1673-11.2011 Jenuwein, T., & Allis, C. D. (2001). Translating the histone code. Science, 293(5532), 1074-1080. doi: 10.1126/science.1063127 Jones, P. A. (2012). Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet, 13(7), 484-492. doi: 10.1038/nrg3230 Katada, S., Imhof, A., & Sassone-Corsi, P. (2012). Connecting threads: epigenetics and metabolism. Cell, 148(1-2), 24-28. doi: 10.1016/j.cell.2012.01.001 Kuzumaki, N., Ikegami, D., Tamura, R., Hareyama, N., Imai, S., Narita, M., . . . Suzuki, T. (2011). Hippocampal epigenetic modification at the brain-derived neurotrophic factor gene induced by an enriched environment. Hippocampus, 21(2), 127-132. doi: 10.1002/hipo.20775 Labonte, B., Suderman, M., Maussion, G., Navaro, L., Yerko, V., Mahar, I., . . . Turecki, G. (2012). Genome-wide epigenetic regulation by early-life trauma. Arch Gen Psychiatry, 69(7), 722-731. doi: 10.1001/archgenpsychiatry.2011.2287 Liu, D., Diorio, J., Tannenbaum, B., Caldji, C., Francis, D., Freedman, A., . . . Meaney, M. J. (1997). Maternal care, hippocampal glucocorticoid receptors, and hypothalamic-pituitary-adrenal responses to stress. Science, 277(5332), 1659-1662. Lu, Y., Christian, K., & Lu, B. (2008). BDNF: a key regulator for protein synthesis-dependent LTP and long-term memory? Neurobiol Learn Mem, 89(3), 312-323. doi: 10.1016/j.nlm.2007.08.018 Minichiello, L. (2009). TrkB signalling pathways in LTP and learning. Nat Rev Neurosci, 10(12), 850-860. doi: 10.1038/nrn2738 Morley-Fletcher, S., Rea, M., Maccari, S., & Laviola, G. (2003). Environmental enrichment during adolescence reverses the effects of prenatal stress on play behaviour and HPA axis reactivity in rats. Eur J Neurosci, 18(12), 3367-3374. Mueller, B. R., & Bale, T. L. (2008). Sex-specific programming of offspring emotionality after stress early in pregnancy. J Neurosci, 28(36), 9055-9065. doi: 10.1523/jneurosci.1424-08.2008 Murgatroyd, C., Patchev, A. V., Wu, Y., Micale, V., Bockmuhl, Y., Fischer, D., . . . Spengler, D. (2009). Dynamic DNA methylation programs persistent adverse effects of early-life stress. Nat Neurosci, 12(12), 1559-1566. doi: 10.1038/nn.2436 Musumeci, G., & Minichiello, L. (2011). BDNF-TrkB signalling in fear learning: from genetics to neural networks. Rev Neurosci, 22(3), 303-315. doi: 10.1515/rns.2011.031 Mychasiuk, R., Zahir, S., Schmold, N., Ilnytskyy, S., Kovalchuk, O., & Gibb, R. (2012). Parental enrichment and offspring development: modifications to brain, behavior and the epigenome. Behav Brain Res, 228(2), 294-298. doi: 10.1016/j.bbr.2011.11.036 Nasca, C., Xenos, D., Barone, Y., Caruso, A., Scaccianoce, S., Matrisciano, F., . . . Nicoletti, F. (2013). L-acetylcarnitine causes rapid antidepressant effects through the epigenetic induction of mGlu2 receptors. Proc Natl Acad Sci U S A, 110(12), 4804-4809. doi: 10.1073/pnas.1216100110 Naumova, O. Y., Lee, M., Koposov, R., Szyf, M., Dozier, M., & Grigorenko, E. L. (2012). Differential patterns of whole-genome DNA methylation in institutionalized children and children raised by their biological parents. Dev Psychopathol, 24(1), 143-155. doi: 10.1017/s0954579411000605 Nithianantharajah, J., & Hannan, A. J. (2006). Enriched environments, experience-dependent plasticity and disorders of the nervous system. Nat Rev Neurosci, 7(9), 697-709. doi: 10.1038/nrn1970 Oberlander, T. F., Weinberg, J., Papsdorf, M., Grunau, R., Misri, S., & Devlin, A. M. (2008). Prenatal exposure to maternal depression, neonatal methylation of human glucocorticoid receptor gene (NR3C1) and infant cortisol stress responses. Epigenetics, 3(2), 97-106. Ooi, S. K., Qiu, C., Bernstein, E., Li, K., Jia, D., Yang, Z., . . . Bestor, T. H. (2007). DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature, 448(7154), 714-717. doi: 10.1038/nature05987 Pariante, C. M., & Lightman, S. L. (2008). The HPA axis in major depression: classical theories and new developments. Trends Neurosci, 31(9), 464-468. doi: 10.1016/j.tins.2008.06.006 Roth, T. L., Lubin, F. D., Funk, A. J., & Sweatt, J. D. (2009). Lasting epigenetic influence of early-life adversity on the BDNF gene. Biol Psychiatry, 65(9), 760-769. Roth, T. L., Zoladz, P. R., Sweatt, J. D., & Diamond, D. M. (2011). Epigenetic modification of hippocampal Bdnf DNA in adult rats in an animal model of post-traumatic stress disorder. J Psychiatr Res, 45(7), 919-926. doi: 10.1016/j.jpsychires.2011.01.013 Sapolsky, R. M., Meaney, M. J., & McEwen, B. S. (1985). The development of the glucocorticoid receptor system in the rat limbic brain. III. Negative-feedback regulation. Brain Res, 350(1-2), 169-173. Sato, F., Tsuchiya, S., Meltzer, S. J., & Shimizu, K. (2011). MicroRNAs and epigenetics. FEBS J, 278(10), 1598-1609. doi: 10.1111/j.1742-4658.2011.08089.x Schlinzig, T., Johansson, S., Gunnar, A., Ekstrom, T. J., & Norman, M. (2009). Epigenetic modulation at birth – altered DNA-methylation in white blood cells after Caesarean section. Acta Paediatr, 98(7), 1096-1099. doi: 10.1111/j.1651-2227.2009.01371.x Suri, D., Veenit, V., Sarkar, A., Thiagarajan, D., Kumar, A., Nestler, E. J., . . . Vaidya, V. A. (2012). Early Stress Evokes Age-Dependent Biphasic Changes in Hippocampal Neurogenesis, Bdnf Expression, and Cognition. Biol Psychiatry. doi: 10.1016/j.biopsych.2012.10.023 Szulwach, K. E., Li, X., Li, Y., Song, C. X., Wu, H., Dai, Q., . . . Jin, P. (2011). 5-hmC-mediated epigenetic dynamics during postnatal neurodevelopment and aging. Nat Neurosci, 14(12), 1607-1616. doi: 10.1038/nn.2959 Tsankova, N., Renthal, W., Kumar, A., & Nestler, E. J. (2007). Epigenetic regulation in psychiatric disorders. Nat Rev Neurosci, 8(5), 355-367. doi: 10.1038/nrn2132 Tsankova, N. M., Berton, O., Renthal, W., Kumar, A., Neve, R. L., & Nestler, E. J. (2006). Sustained hippocampal chromatin regulation in a mouse model of depression and antidepressant action. Nat Neurosci, 9(4), 519-525. doi: 10.1038/nn1659 Uddin, M., Aiello, A. E., Wildman, D. E., Koenen, K. C., Pawelec, G., de Los Santos, R., . . . Galea, S. (2010). Epigenetic and immune function profiles associated with posttraumatic stress disorder. Proc Natl Acad Sci U S A, 107(20), 9470-9475. doi: 10.1073/pnas.0910794107 van der Horst, F. C., & van der Veer, R. (2008). Loneliness in infancy: Harry Harlow, John Bowlby and issues of separation. Integr Psychol Behav Sci, 42(4), 325-335. doi: 10.1007/s12124-008-9071-x Weaver, I. C., Cervoni, N., Champagne, F. A., D’Alessio, A. C., Sharma, S., Seckl, J. R., . . . Meaney, M. J. (2004). Epigenetic programming by maternal behavior. Nat Neurosci, 7(8), 847-854. doi: 10.1038/nn1276 Weaver, I. C., Champagne, F. A., Brown, S. E., Dymov, S., Sharma, S., Meaney, M. J., & Szyf, M. (2005). Reversal of maternal programming of stress responses in adult offspring through methyl supplementation: altering epigenetic marking later in life. J Neurosci, 25(47), 11045-11054. doi: 10.1523/jneurosci.3652-05.2005 Weaver, I. C., Diorio, J., Seckl, J. R., Szyf, M., & Meaney, M. J. (2004). Early environmental regulation of hippocampal glucocorticoid receptor gene expression: characterization of intracellular mediators and potential genomic target sites. Ann N Y Acad Sci, 1024, 182-212. doi: 10.1196/annals.1321.099 Weaver, I. C., Meaney, M. J., & Szyf, M. (2006). Maternal care effects on the hippocampal transcriptome and anxiety-mediated behaviors in the offspring that are reversible in adulthood. Proc Natl Acad Sci U S A, 103(9), 3480-3485. doi: 10.1073/pnas.0507526103 Whittle, N., Schmuckermair, C., Gunduz Cinar, O., Hauschild, M., Ferraguti, F., Holmes, A., & Singewald, N. (2013). Deep brain stimulation, histone deacetylase inhibitors and glutamatergic drugs rescue resistance to fear extinction in a genetic mouse model. Neuropharmacology, 64, 414-423. doi: 10.1016/j.neuropharm.2012.06.001 Yehuda, R., Bell, A., Bierer, L. M., & Schmeidler, J. (2008). Maternal, not paternal, PTSD is related to increased risk for PTSD in offspring of Holocaust survivors. J Psychiatr Res, 42(13), 1104-1111. doi: 10.1016/j.jpsychires.2008.01.002 Yehuda, R., & Bierer, L. M. (2009). The relevance of epigenetics to PTSD: implications for the DSM-V. J Trauma Stress, 22(5), 427-434. doi: 10.1002/jts.20448